

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0070.9
(437 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
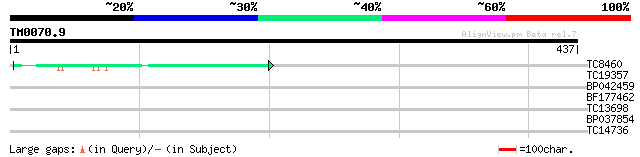
TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F... 50 7e-07
TC19357 weakly similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5... 37 0.008
BP042459 28 2.3
BF177462 28 3.0
TC13698 similar to GB|AAO63291.1|28950735|BT005227 At4g20000 {Ar... 27 5.1
BP037854 27 6.6
TC14736 similar to UP|Q9LHS4 (Q9LHS4) Human RAN binding protein ... 27 6.6
>TC8460 similar to GB|AAN73294.1|25141199|BT002297 At5g12010/F14F18_180
{Arabidopsis thaliana;}, partial (36%)
Length = 880
Score = 50.1 bits (118), Expect = 7e-07
Identities = 58/228 (25%), Positives = 91/228 (39%), Gaps = 28/228 (12%)
Frame = +3
Query: 4 NNLPDWNTFYNECVEDFMNDTFVEDMMQQEMEF-----YQQ---------QQHANTVRPK 49
NN P+WN +NDT + D++ + F +QQ Q H P+
Sbjct: 99 NNDPEWN----------LNDTVLNDLLTSFIHFDDTKPHQQLMDPNPIANQSHIEQFEPE 248
Query: 50 KTRRVIKRDREAGN------ERLM-----KDYFSK--NPVYTEELFRRRFRMRKHVFLRI 96
K+ R + RL KD++ + +P + EE FRR FRM K F I
Sbjct: 249 FDEPPTKKPRRSDGTATTPPRRLWVKDRSKDWWERCSHPDFPEEEFRRSFRMSKATFEMI 428
Query: 97 VEALGSYNPYFLMSVDAVGRQGLSPLQKCTAAIRMLAYGSPADSVDEYVRIGESTAIECL 156
L S + + + R+ + Q+ I LA G P V + +G ST + +
Sbjct: 429 CRELDS----AVTKKNTMLREAIPVRQRVAVCIWRLATGDPLRLVAKRFGLGISTCHKLV 596
Query: 157 KNFVEGVCEVFGGQYLRRPNEEDMTRLLQWGES-RGFPGMLGSIDCMH 203
+ V ++LR P+E MT E+ G P + G++ H
Sbjct: 597 LEVCSAIKTVLMPKFLRWPDEAAMTAAKSEFEALSGIPNIGGAMYTTH 740
>TC19357 weakly similar to UP|Q9FHY5 (Q9FHY5) Emb|CAB72466.1 (At5g41980),
partial (16%)
Length = 577
Score = 36.6 bits (83), Expect = 0.008
Identities = 30/118 (25%), Positives = 57/118 (47%)
Frame = +2
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDER 370
K+ F R R + R+F VL++RF I+ P+ + + I+ A +LHN+I ++R
Sbjct: 5 KELFNHRHYFLRGAILRSFTVLKARFPILI-PAPQYSFQIQRDIVIAACVLHNIIRREDR 181
Query: 371 NTYKGNFVYEQVNNDISDAEVLSGPIPAFRNMLERRAHQIEKSIHRQLQADLVEHIWD 428
N + N + ++++ + L + +M E+ A + SI + D + WD
Sbjct: 182 NDWLFNSLGGTPVDELTVLDELP-DVQLLSSMQEQLAFSLRDSIASAMWDDFLNK-WD 349
>BP042459
Length = 529
Score = 28.5 bits (62), Expect = 2.3
Identities = 20/64 (31%), Positives = 33/64 (51%), Gaps = 1/64 (1%)
Frame = -1
Query: 114 VGRQGLSPLQKCTAAIRMLAYGSPADSVDEYVRIGESTAIECLKNFVEGVCEVF-GGQYL 172
+GR G + R+ +Y S D Y+ +G++ ++ + V G EV+ GGQYL
Sbjct: 358 IGRNGFEETYLYSPGSRVESY-----SKDAYICVGQAAVLQPI---VLGPEEVWKGGQYL 203
Query: 173 RRPN 176
+ PN
Sbjct: 202 QNPN 191
>BF177462
Length = 417
Score = 28.1 bits (61), Expect = 3.0
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 6/51 (11%)
Frame = -2
Query: 318 QEAARKDVERAF------GVLQSRFAIVRGPSRWWHPNDMKSIIYACIILH 362
+ AA + V R F G+ S FAI+ G WW N S+I +ILH
Sbjct: 194 ERAAEEGVGREFQGR*ERGIGGSEFAILEGWRVWWSLN--PSLIQVLLILH 48
>TC13698 similar to GB|AAO63291.1|28950735|BT005227 At4g20000 {Arabidopsis
thaliana;}, partial (10%)
Length = 419
Score = 27.3 bits (59), Expect = 5.1
Identities = 10/15 (66%), Positives = 12/15 (79%)
Frame = +2
Query: 30 MQQEMEFYQQQQHAN 44
MQ +M+FYQQQ H N
Sbjct: 77 MQMQMQFYQQQPHMN 121
>BP037854
Length = 462
Score = 26.9 bits (58), Expect = 6.6
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = -1
Query: 227 IMLEAVASQDLWIWHAFFGIA 247
++ +V + DLW W FFG A
Sbjct: 282 LIFSSVLALDLWFWELFFGTA 220
>TC14736 similar to UP|Q9LHS4 (Q9LHS4) Human RAN binding protein 16-like,
partial (5%)
Length = 757
Score = 26.9 bits (58), Expect = 6.6
Identities = 10/23 (43%), Positives = 13/23 (56%)
Frame = +2
Query: 329 FGVLQSRFAIVRGPSRWWHPNDM 351
FGV F I+ P WWHP ++
Sbjct: 185 FGVCILSFWIISFPPIWWHPGNL 253
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,233,215
Number of Sequences: 28460
Number of extensions: 94626
Number of successful extensions: 512
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 510
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 512
length of query: 437
length of database: 4,897,600
effective HSP length: 93
effective length of query: 344
effective length of database: 2,250,820
effective search space: 774282080
effective search space used: 774282080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0070.9


