

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
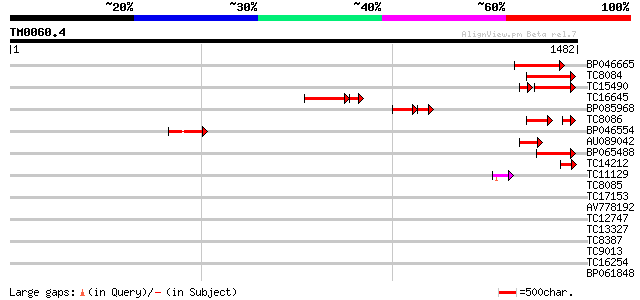
Query= TM0060.4
(1482 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BP046665 161 8e-40
TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombin... 156 2e-38
TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like prote... 132 4e-37
TC16645 132 9e-37
BP085968 81 2e-25
TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination p... 82 4e-22
BP046554 85 7e-17
AU089042 85 1e-16
BP065488 82 5e-16
TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%) 57 2e-08
TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protei... 48 1e-05
TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meioti... 33 0.33
TC17153 UP|AAQ87023 (AAQ87023) VDAC3.1, complete 31 1.3
AV778192 30 2.1
TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-l... 30 3.6
TC13327 similar to UP|O48939 (O48939) Polyphosphoinositide bindi... 29 4.8
TC8387 similar to UP|BCA2_ARATH (Q9M439) Branched-chain amino ac... 29 6.2
TC9013 29 6.2
TC16254 similar to UP|O04252 (O04252) T10M13.13 (CTP synthase-li... 28 8.1
BP061848 28 8.1
>BP046665
Length = 524
Score = 161 bits (407), Expect = 8e-40
Identities = 76/131 (58%), Positives = 101/131 (77%), Gaps = 1/131 (0%)
Frame = -1
Query: 1320 DIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGR 1379
DI C GIP+HK+ LKEGAP+ML+RN+ + G CNGTR++V L NV+ VI+ T++G
Sbjct: 524 DITCFGIPNHKITLKEGAPIMLLRNIYQAVGFCNGTRLIVADLGTNVIKATVIT*TNIGD 345
Query: 1380 QVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNPVFSHGQL 1439
+FI RMD++P+D P KF+RRQFP+ L AMTINKSQG++LSHVGLYL VF+HGQL
Sbjct: 344 DIFIPRMDMVPSDSGYPFKFERRQFPISLCSAMTINKSQGQSLSHVGLYLSRHVFTHGQL 165
Query: 1440 YVAI-SRVKTR 1449
YVA+ +++K R
Sbjct: 164 YVALHAKIKKR 132
Score = 32.3 bits (72), Expect = 0.56
Identities = 14/45 (31%), Positives = 28/45 (62%)
Frame = -2
Query: 1428 YLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVF 1472
Y+ ++SH +S +++R GLK+L+ +E+ + TKN+V+
Sbjct: 202 YIFLAMYSHMVSCTLLSMLRSRKGLKLLVLDEEEKVTNTTKNVVY 68
>TC8084 similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (4%)
Length = 560
Score = 156 bits (395), Expect = 2e-38
Identities = 72/129 (55%), Positives = 100/129 (76%)
Frame = +1
Query: 1351 LCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSF 1410
LCNGTR++V L V+ VI+GT++G +FI R+D++P+D P KF+RR FP+ L F
Sbjct: 61 LCNGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCF 240
Query: 1411 AMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNI 1470
AMTINKSQG++LSHV LYL PVF+HGQLYVA+SRV++R GLK+L+ +E+ + TKN+
Sbjct: 241 AMTINKSQGQSLSHVSLYLSRPVFTHGQLYVALSRVRSRKGLKLLVLDEEEKVTNTTKNV 420
Query: 1471 VFKEVFQRI 1479
V++EVF+ I
Sbjct: 421 VYREVFENI 447
>TC15490 weakly similar to UP|Q9LTU4 (Q9LTU4) Helicase-like protein, partial
(8%)
Length = 634
Score = 132 bits (332), Expect(2) = 4e-37
Identities = 61/109 (55%), Positives = 85/109 (77%)
Frame = +2
Query: 1371 VISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLP 1430
VI+GTH+G + I RMD++P+D S P KF+RRQ P+ L FAMTINKSQG++LSHVGLYL
Sbjct: 149 VITGTHIGDDISIPRMDMVPSDSSYPFKFERRQSPISLCFAMTINKSQGRSLSHVGLYLS 328
Query: 1431 NPVFSHGQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
PV +HG LYVA+ RV++R LK+L+ +E+ + TKN+V++E+F+ I
Sbjct: 329 RPVSTHG*LYVALPRVRSRK*LKLLVLDEEEKMTNTTKNVVYREIFENI 475
Score = 41.2 bits (95), Expect(2) = 4e-37
Identities = 18/32 (56%), Positives = 25/32 (77%)
Frame = +1
Query: 1334 KEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPN 1365
KEGA +ML+RN+ ++GLCNGTR++V L N
Sbjct: 43 KEGALIMLLRNIVQASGLCNGTRLIVVDLGIN 138
>TC16645
Length = 596
Score = 132 bits (331), Expect(2) = 9e-37
Identities = 60/119 (50%), Positives = 84/119 (70%)
Frame = +1
Query: 770 MEANCKYPLGRGLTYAEFPSSFVYDKKKKEWHPRKKGICIGRMNFVPPGSGELYYLRMLL 829
M AN KY GR LTYAEFPS F+Y + +W P++K +GR+ F+ PG GE YY+R+LL
Sbjct: 7 MNANKKYQEGRNLTYAEFPSEFLYHQHTTKWVPKQKRFSLGRLPFIAPGMGENYYMRVLL 186
Query: 830 NVQRGCTSFEDLRSVNNHVHDTFRQACLALSLLKDDREYIDGILDVATFASGSYIRELF 888
+Q+GC SF+ +++V V+ TF AC A+ LL+DDREY+DGI + F SG+ +R+LF
Sbjct: 187 TMQKGCDSFKSIKTVKGVVYPTFHDACEAMGLLEDDREYVDGISISSEFGSGTQLRKLF 363
Score = 40.4 bits (93), Expect(2) = 9e-37
Identities = 16/37 (43%), Positives = 25/37 (67%)
Frame = +2
Query: 888 FVSFLLGNAMADPSHVWRETWSVLADGIVHALRRTHN 924
F L+ N ++ P VWR++WS+L DGI++ R+T N
Sbjct: 362 FTRMLMTNLISRPQEVWRKSWSLLCDGILYDRRKTLN 472
>BP085968
Length = 341
Score = 81.3 bits (199), Expect(2) = 2e-25
Identities = 38/65 (58%), Positives = 53/65 (81%)
Frame = +2
Query: 1002 QAKIYEEIISAVNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLL 1061
Q+K+Y++I+S+V SD G F F+YG+GG+GKTF+W T + LRS+ ++LNVASSGIAS L
Sbjct: 20 QSKVYKQIMSSVWSDDGGFYFLYGFGGSGKTFVWNTWSSGLRSQGLMVLNVASSGIASWL 199
Query: 1062 LHGGR 1066
L GG+
Sbjct: 200 LPGGK 214
Score = 53.1 bits (126), Expect(2) = 2e-25
Identities = 26/43 (60%), Positives = 32/43 (73%)
Frame = +3
Query: 1066 RTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSLIIWDEAP 1108
RTAHS F I ++ ++ S C I QGS KAELL+ +S IIWDEAP
Sbjct: 213 RTAHSRFSIHISINDISTCNIKQGSQKAELLQKASSIIWDEAP 341
>TC8086 similar to UP|DMC1_HUMAN (Q14565) Meiotic recombination protein
DMC1/LIM15 homolog, partial (12%)
Length = 628
Score = 82.0 bits (201), Expect(2) = 4e-22
Identities = 37/69 (53%), Positives = 51/69 (73%)
Frame = +1
Query: 1351 LCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSF 1410
LC+GTR++V L V+ VI+GT++G +FI R+D++P+D P KF+RR FP+ L F
Sbjct: 169 LCHGTRLIVVDLGTYVIKATVITGTNIGDDIFIPRLDMVPSDSGYPFKFERR*FPISLCF 348
Query: 1411 AMTINKSQG 1419
AMTINKSQG
Sbjct: 349 AMTINKSQG 375
Score = 41.2 bits (95), Expect(2) = 4e-22
Identities = 17/36 (47%), Positives = 28/36 (77%)
Frame = +3
Query: 1444 SRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
SRV++R GLK+L+ +E+ + TKN+V++EVF+ I
Sbjct: 369 SRVRSRKGLKLLVLDEEEKVTNTTKNVVYREVFENI 476
>BP046554
Length = 556
Score = 85.1 bits (209), Expect = 7e-17
Identities = 47/105 (44%), Positives = 65/105 (61%), Gaps = 2/105 (1%)
Frame = -3
Query: 414 EEAIHRGDTDLSMVGSRIVLPSSFTGGRRY--MFNNCQDAMGICREYGYPDLFLTMTCNP 471
EEAI +G+T+ S VGS + P + + Q I +++GYPDLF+T TCN
Sbjct: 392 EEAIDKGETNPSSVGSELFCPHLLLVDIAICSIISRMQWPC-IFKKFGYPDLFITFTCNS 216
Query: 472 KWPEIERHVSARCLSAYDRPDLACRVFRMKLDQLMKNLKKGKFFG 516
W EI+R V R L+ D P++ RVF+MKLD+L+ +LKKGK FG
Sbjct: 215 AWCEIQRFVQPRNLNVEDCPNICVRVFKMKLDRLISDLKKGKIFG 81
>AU089042
Length = 191
Score = 84.7 bits (208), Expect = 1e-16
Identities = 37/61 (60%), Positives = 48/61 (78%)
Frame = +3
Query: 1332 VLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGTHVGRQVFIGRMDLMPT 1391
VLK G PVMLM NL +STGLCNGTR++V +L PNV+G ++SGTH+G V+I M+L P+
Sbjct: 6 VLKXGVPVMLMXNLXISTGLCNGTRLIVDYLGPNVIGATILSGTHIGXVVYISMMNLXPS 185
Query: 1392 D 1392
D
Sbjct: 186 D 188
>BP065488
Length = 439
Score = 82.4 bits (202), Expect = 5e-16
Identities = 47/104 (45%), Positives = 67/104 (64%), Gaps = 2/104 (1%)
Frame = -1
Query: 1378 GRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKT-LSHVGLYLPNPVFSH 1436
G +FI RM+++P+ P+KF+R QFP+ L FAMTINKSQ V L +SH
Sbjct: 379 GDDIFIPRMNMVPSVSGDPLKFERCQFPISLCFAMTINKSQXSVHYPTVALISFLACYSH 200
Query: 1437 -GQLYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRI 1479
+LYVA+S V++R GLK+L+ E+ + TKN V++EVF+ I
Sbjct: 199 MDKLYVALSGVRSRKGLKLLVLGEEEKVTNATKNEVYQEVFKNI 68
>TC14212 similar to UP|Q9FN61 (Q9FN61) Gb|AAF07369.1, partial (10%)
Length = 613
Score = 57.4 bits (137), Expect = 2e-08
Identities = 28/43 (65%), Positives = 35/43 (81%)
Frame = +2
Query: 1439 LYVAISRVKTRAGLKILICNEDISQRDVTKNIVFKEVFQRIYS 1481
LYVA+SRVK++ GLKILI ++ S TKNIV+KEVFQ+IYS
Sbjct: 2 LYVAVSRVKSKDGLKILISSDGTSTPGATKNIVYKEVFQKIYS 130
>TC11129 weakly similar to UP|PDA6_MEDSA (P38661) Probable protein disulfide
isomerase A6 precursor (P5) , partial (7%)
Length = 600
Score = 47.8 bits (112), Expect = 1e-05
Identities = 23/62 (37%), Positives = 36/62 (57%), Gaps = 7/62 (11%)
Frame = +2
Query: 1261 YYEDKAILA-------PTLDAVDLVNQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWI 1313
YY D +++ PTL++V+ VN+++L L PGN YLSS + ED +++ W
Sbjct: 410 YYADGGVISLVMY*IFPTLESVEKVNEFMLDLLPGNTTEYLSSDTTCKYDEDTELQS*WF 589
Query: 1314 TT 1315
TT
Sbjct: 590 TT 595
>TC8085 similar to GB|BAB03033.1|11994717|AP001300 AtDMC1 (meiotic
recombination protein)-like protein {Arabidopsis
thaliana;} , partial (8%)
Length = 500
Score = 33.1 bits (74), Expect = 0.33
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +1
Query: 897 MADPSHVWRETWSVLADGIVHALRRT 922
M +P HVW TW +LA+ I H R T
Sbjct: 115 MTNPDHVWNITWKLLANDIQHEYRGT 192
>TC17153 UP|AAQ87023 (AAQ87023) VDAC3.1, complete
Length = 1175
Score = 31.2 bits (69), Expect = 1.3
Identities = 15/47 (31%), Positives = 26/47 (54%)
Frame = -2
Query: 699 DEIKQYHDCRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVIL 745
++I Q H CRY+ C +F+IH K P ++K+ Y + ++L
Sbjct: 406 EDIIQSHFCRYIRVC------INFHIHSKIPTLVKRIYVTNENLILL 284
>AV778192
Length = 399
Score = 30.4 bits (67), Expect = 2.1
Identities = 14/34 (41%), Positives = 22/34 (64%)
Frame = -2
Query: 1165 VMATINSSRLWRFCKVLTLTENMRLFSNSETSDV 1198
+++TINSS + FC +L L R F S+TS++
Sbjct: 131 LLSTINSSNYFLFCSLLLLCILFRRFIESDTSNI 30
>TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-like protein,
partial (43%)
Length = 624
Score = 29.6 bits (65), Expect = 3.6
Identities = 16/41 (39%), Positives = 21/41 (51%)
Frame = -2
Query: 1061 LLHGGRTAHSLFCIPLNADEDSCCGIVQGSPKAELLKLSSL 1101
L G + H+L C + C I+ G PK E LKLS+L
Sbjct: 332 LCEGIQAVHALLCYEF----ERTCNILDGFPKKESLKLSNL 222
>TC13327 similar to UP|O48939 (O48939) Polyphosphoinositide binding protein
Ssh1p, partial (19%)
Length = 525
Score = 29.3 bits (64), Expect = 4.8
Identities = 20/89 (22%), Positives = 44/89 (48%), Gaps = 6/89 (6%)
Frame = +3
Query: 916 VHALRRTHNNLELVIDNE----HLKQL--CLMEIEKLLMINGRSLKDFDNMPCVDSNILI 969
+H ++H N + ++N+ HL Q C++ + + ++ R+L N PC+
Sbjct: 177 LHKFEKSHGNDDR*VNNQWYLCHLIQFQTCVVSV---VYVSLRNLGSSHNKPCIVPQCKR 347
Query: 970 QYGNILLFNELNFDTVEMSKLHVECLNKL 998
+ NI+ +++++ ++ L CLN L
Sbjct: 348 EMSNIIFYHQVSSLSLIFCFLSS*CLNNL 434
>TC8387 similar to UP|BCA2_ARATH (Q9M439) Branched-chain amino acid
aminotransferase 2, chloroplast precursor (Atbcat-2) ,
partial (84%)
Length = 1883
Score = 28.9 bits (63), Expect = 6.2
Identities = 19/79 (24%), Positives = 37/79 (46%)
Frame = +2
Query: 256 VALRLFRHRVNDPKTYNLPTVDEVAALIVEDFDTSDCGRDIILRTSSGNLQRIYDTHSSF 315
V+L F + PKT+ +PT ++++ C R ++LR + T++S+
Sbjct: 35 VSLHYFHRFIAYPKTHKIPT----NYMMIQRTSQFPCLRKLLLRAGYPSSSSKMGTYNSY 202
Query: 316 LPLQYPLIFPYGEEGFSDE 334
PL P ++ +SD+
Sbjct: 203 ASQPSPLEIP--DQSYSDD 253
>TC9013
Length = 766
Score = 28.9 bits (63), Expect = 6.2
Identities = 12/26 (46%), Positives = 18/26 (69%)
Frame = +3
Query: 1454 ILICNEDISQRDVTKNIVFKEVFQRI 1479
ILIC+ D S + T N+V+ EVF+ +
Sbjct: 627 ILICDGDDSNSNSTSNVVYTEVFRTV 704
>TC16254 similar to UP|O04252 (O04252) T10M13.13 (CTP synthase-like protein)
, partial (7%)
Length = 515
Score = 28.5 bits (62), Expect = 8.1
Identities = 25/79 (31%), Positives = 35/79 (43%), Gaps = 2/79 (2%)
Frame = +1
Query: 66 YFDLGEMNMACQYCGAILWYHERAQKAKNAISPDFSICCMKGKISLPYLQDA--PTLLSN 123
YF E + CQ+ ++ + E K F C KGK L L DA + L+
Sbjct: 148 YFQ-SENDHPCQFYSSVTYSSEDCHGGKICWFNLFICICCKGKPLLGTLLDAYRSSFLAR 324
Query: 124 LLTNIDPRSGHFIDNIRSY 142
+IDPR FI ++R Y
Sbjct: 325 TFHDIDPRV-VFIAHLREY 378
>BP061848
Length = 379
Score = 28.5 bits (62), Expect = 8.1
Identities = 18/61 (29%), Positives = 29/61 (47%)
Frame = +3
Query: 126 TNIDPRSGHFIDNIRSYNSMFAFTSIGGKVDGSVNNGQGPPQFVISGQNYHRIGSLLPAE 185
TNIDP+ G+ ID++ S + ++ S NG+G ++G+ RI P
Sbjct: 204 TNIDPQFGNIIDSLHSSEKTVSKHYPSVSLETSSENGRGS---TVAGE*EFRINGKGPVA 374
Query: 186 G 186
G
Sbjct: 375 G 377
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,585,439
Number of Sequences: 28460
Number of extensions: 385607
Number of successful extensions: 1632
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 1620
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1630
length of query: 1482
length of database: 4,897,600
effective HSP length: 102
effective length of query: 1380
effective length of database: 1,994,680
effective search space: 2752658400
effective search space used: 2752658400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0060.4


