

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0059.7
(1706 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
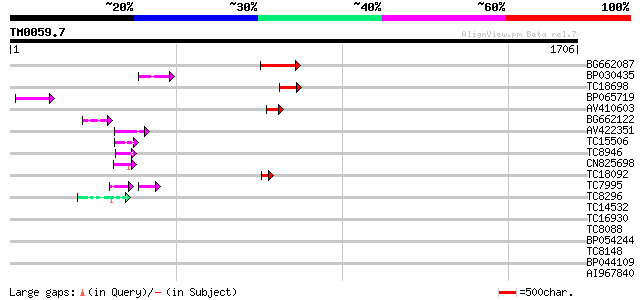
Score E
Sequences producing significant alignments: (bits) Value
BG662087 168 8e-42
BP030435 62 8e-10
TC18698 59 6e-09
BP065719 57 2e-08
AV410603 50 3e-06
BG662122 49 7e-06
AV422351 47 3e-05
TC15506 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich prote... 47 3e-05
TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine b... 45 1e-04
CN825698 45 1e-04
TC18092 44 2e-04
TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferas... 35 4e-04
TC8296 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1... 42 6e-04
TC14532 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor ... 42 0.001
TC16930 39 0.007
TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like arabinogalac... 39 0.007
BP054244 39 0.007
TC8148 homologue to PIR|E86216|E86216 protein T23G18.10 [importe... 39 0.009
BP044109 39 0.009
AI967840 38 0.012
>BG662087
Length = 373
Score = 168 bits (425), Expect = 8e-42
Identities = 78/120 (65%), Positives = 96/120 (80%)
Frame = +1
Query: 754 NGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEE 813
+GKWRM VDYTDLNKACPKD +PLPSID LVD +S E LSLMDAYSGY+QI MH DE+
Sbjct: 13 SGKWRMWVDYTDLNKACPKDSYPLPSIDKLVDGASDNELLSLMDAYSGYHQIKMHPSDED 192
Query: 814 KTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDH 873
KTAF+T R YCY +PFGLKNA ATYQ +M R+F D +G+++EVY+D++IVK+ +H
Sbjct: 193 KTAFMTARVNYCYQTIPFGLKNAGATYQXLMDRVFXDXVGRNMEVYLDNMIVKSALRANH 372
>BP030435
Length = 533
Score = 62.0 bits (149), Expect = 8e-10
Identities = 39/106 (36%), Positives = 57/106 (52%)
Frame = +1
Query: 389 VGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPRQKVAITFTEDDYGEDTGEE 448
V +IAGG+ GG T+ ++KR R + + G HP ITF+ DD+
Sbjct: 232 VHTIAGGFGGGGITSAARKRYARAVNTVT-EVLFGFSHP-----VITFSSDDFIRIKPHL 393
Query: 449 DDPIVIEALIGNGKIRRTLIDTGSSADIMFYDAYKSLGLSVKDLLP 494
D+PIVI + ++R L+D GSSADI++ + LGL+ DL P
Sbjct: 394 DNPIVILLRVNQLNVQRVLLDQGSSADIIYGGVFDRLGLNEADLTP 531
>TC18698
Length = 808
Score = 58.9 bits (141), Expect = 6e-09
Identities = 29/67 (43%), Positives = 44/67 (65%)
Frame = -2
Query: 812 EEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGG 871
++KT +R Y Y ++P GLKN TYQR+M +IF + K+VEVY++D+IVK+ +
Sbjct: 807 KKKTTLKINRVNYYYQVMPLGLKNI*TTYQRLMDKIFHKQI*KNVEVYVEDMIVKSSQE* 628
Query: 872 DHAADLA 878
H DL+
Sbjct: 627 FHRGDLS 607
>BP065719
Length = 567
Score = 57.4 bits (137), Expect = 2e-08
Identities = 36/117 (30%), Positives = 53/117 (44%), Gaps = 2/117 (1%)
Frame = +3
Query: 19 EIMAFPFPQGWARPKVKLYEGDSDPQ--EHVNFFVGAMQYAGASDPLYCRCFPMSLGKGP 76
E++ P+GW PK + GDS EH+ + ++ L + FP SL K
Sbjct: 144 EVLESELPRGWKVPKFTKFSGDSGESTVEHIARYQIEAGDLAINENLKMKYFPSSLTKNA 323
Query: 77 MNWFQNLPSNSLHDWNGVVTCFLAQYSSVRSIPKTAQTLALVKQKEKESLKAFLNRF 133
WF L S+H W + F Q+ + + LA VK+K ES+ +LNRF
Sbjct: 324 FTWFTTLAPRSVHTWAQLERIFHEQFFR-GECKVSXKDLASVKRKPAESIDDYLNRF 491
>AV410603
Length = 162
Score = 50.1 bits (118), Expect = 3e-06
Identities = 22/51 (43%), Positives = 32/51 (62%)
Frame = +1
Query: 772 KDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRG 822
KD FP+P++D L+D G ++ S +D SGY+QI + +D KT F T G
Sbjct: 10 KDSFPMPTVDELLDELRGSQFFSKLDLRSGYHQILVKPEDRHKTVFRTHHG 162
>BG662122
Length = 386
Score = 48.9 bits (115), Expect = 7e-06
Identities = 33/100 (33%), Positives = 56/100 (56%), Gaps = 11/100 (11%)
Frame = +2
Query: 220 RRKGQDERKP------KRKKYDSYTPLNSSLSRILREKASTDLRDR-----PPPLLTRGD 268
RR+ D R+P +R++ D LN+ L+ IL++ + + + PPP RG
Sbjct: 29 RREALDRRRP*QPMGSRRREEDM--ELNAHLTDILQDVKAAHMVGKSGQS*PPP--RRG- 193
Query: 269 KLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRY 308
+D+ + CE+ S H+ D+C LK ++E+LI+ GRL +Y
Sbjct: 194 -IDTTK*CEYRRSVVHDIDDCFTLKREIEKLIKMGRLKQY 310
>AV422351
Length = 489
Score = 46.6 bits (109), Expect = 3e-05
Identities = 37/109 (33%), Positives = 52/109 (46%), Gaps = 4/109 (3%)
Frame = +3
Query: 316 LPRPRSPPPRRTP-TPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERR--RSPDR 372
LPR R PPP P PP+ R Q ++ ++ R PP+R SP RR R+R+ R+ R
Sbjct: 162 LPRHRPPPPNAIPRPPPQSRHQPHLRQENSRLRRDPPPKRCSPP*RRHPRDRQPHRATFR 341
Query: 373 HGRDEVRRQHGSNIVDVGSIAGGWAAG-GPTNNSQKRSTRVIMSAAGRP 420
G + + D+ GG AAG P ++ S R A+G P
Sbjct: 342 AGGFQPEEKPPR---DLCGAPGGSAAGRSPDQPLEETSPRPSRRASGTP 479
Score = 30.0 bits (66), Expect = 3.2
Identities = 29/76 (38%), Positives = 32/76 (41%), Gaps = 13/76 (17%)
Frame = +2
Query: 307 RYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKT------PPRRRSP--E 358
RY +T P P PP R+ TPPR QT +A T P RR S E
Sbjct: 191 RYPPTTTSISP-PTPPPARKQSTPPRSSAQTMFSALTASSTGSTASSCHVPCRRISTGGE 367
Query: 359 RRRRS-----RERRRS 369
RS RERRRS
Sbjct: 368 ASARSLRCSRRERRRS 415
>TC15506 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein, partial
(24%)
Length = 677
Score = 46.6 bits (109), Expect = 3e-05
Identities = 35/77 (45%), Positives = 41/77 (52%), Gaps = 2/77 (2%)
Frame = +2
Query: 314 GALPRPRSPPPR-RTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERR-RRSRERRRSPD 371
G+ R RSPPP R +PP R ++PP+R RR R RSP RR RS R SP
Sbjct: 23 GSPGRRRSPPPPPRRRSPPPRRARSPPRRSPIGRR-----RSRSPIRRPARSNSRSFSP- 184
Query: 372 RHGRDEVRRQHGSNIVD 388
R GR VRR S+ D
Sbjct: 185 RRGRPPVRRGRSSSFSD 235
Score = 40.0 bits (92), Expect = 0.003
Identities = 34/84 (40%), Positives = 36/84 (42%), Gaps = 25/84 (29%)
Frame = +2
Query: 317 PRPRSPPPRRTPTPPRH-----RDQTPPKRRSAE--------RRDKTPPRR--------- 354
PR RSPPPRR +PPR R P RR A RR + P RR
Sbjct: 59 PRRRSPPPRRARSPPRRSPIGRRRSRSPIRRPARSNSRSFSPRRGRPPVRRGRSSSFSDS 238
Query: 355 ---RSPERRRRSRERRRSPDRHGR 375
R RR RSR RR P R GR
Sbjct: 239 PSPRKVSRRSRSRSPRR-PLRGGR 307
>TC8946 homologue to GB|AAM66972.1|21617922|AY088650 pyridoxine
biosynthesis protein-like {Arabidopsis thaliana;} ,
partial (45%)
Length = 503
Score = 45.1 bits (105), Expect = 1e-04
Identities = 29/68 (42%), Positives = 33/68 (47%), Gaps = 3/68 (4%)
Frame = +3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTP---PRRRSPERRRRSRERRRSPDRHG 374
RPRS PRR R R +P +RRS R P PRR RRRRS ER P H
Sbjct: 165 RPRSDAPRRRHHGRRQRPTSPHRRRSRRLRRHGPRACPRRHPRPRRRRSHER---PSAHQ 335
Query: 375 RDEVRRQH 382
++ R H
Sbjct: 336 GNQTSRHH 359
Score = 33.1 bits (74), Expect = 0.38
Identities = 22/64 (34%), Positives = 25/64 (38%), Gaps = 12/64 (18%)
Frame = +3
Query: 324 PRRTPTPPRHRDQTPPKRRSAERRDKTPPR---------RRSPE---RRRRSRERRRSPD 371
PRR P P R R P + + PR RRSP+ RRR R R R P
Sbjct: 276 PRRHPRPRRRRSHERPSAHQGNQTSRHHPRHGQSPHRPFRRSPDPRSHRRRLRRRERGPH 455
Query: 372 RHGR 375
R
Sbjct: 456 SRRR 467
Score = 30.0 bits (66), Expect = 3.2
Identities = 30/117 (25%), Positives = 48/117 (40%), Gaps = 12/117 (10%)
Frame = +2
Query: 1452 TTPSGLRPSH*QRLPQLEYNASSTRTSSAGSESPES*SQTMGPSLPLRGSETSWMGYTSN 1511
T P RP R P ++S R S+A S S P+ P + + +
Sbjct: 116 TAPQSPRP----RNPPSPSKSASLRCSAAASSWTSS-----TPNKPASPKKPAPAPSWPS 268
Query: 1512 IVSPPSSTPR------------RTDRQSQPTGSFSGASGKDWGAPRATGPNSSITSS 1556
VSPP+S P+ R + P+ S+ + PR++ P++SITS+
Sbjct: 269 SVSPPTSAPKAASLA*ATLSSSRKSNKPSPSPSWPKPASAISSKPRSSKPSASITST 439
>CN825698
Length = 704
Score = 45.1 bits (105), Expect = 1e-04
Identities = 30/85 (35%), Positives = 37/85 (43%), Gaps = 14/85 (16%)
Frame = +1
Query: 312 STGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRD----KTPPRRRS----------P 357
S A P P PPP + PP D PP R +RRD PP RR P
Sbjct: 121 SASAPPPPPPPPPPPSLPPPSDSDLPPPPARRRDRRDDRDFDRPPNRRDYYDRSNNSPPP 300
Query: 358 ERRRRSRERRRSPDRHGRDEVRRQH 382
+ R R +RRRSP + + R+H
Sbjct: 301 KDRDRDFKRRRSPSPPSQRDRDRRH 375
Score = 33.1 bits (74), Expect = 0.38
Identities = 22/58 (37%), Positives = 27/58 (45%), Gaps = 2/58 (3%)
Frame = +1
Query: 321 SPPPR-RTPTPPRHRDQTPPKRRSAERR-DKTPPRRRSPERRRRSRERRRSPDRHGRD 376
SPPP+ R R R +PP +R +RR PP +RS R R DR G D
Sbjct: 289 SPPPKDRDRDFKRRRSPSPPSQRDRDRRHSPPPPYKRSRRGSPRGGYRYGPDDRFGYD 462
>TC18092
Length = 519
Score = 44.3 bits (103), Expect = 2e-04
Identities = 23/35 (65%), Positives = 26/35 (73%)
Frame = -2
Query: 759 MCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYL 793
MCV TD+NK CP+ P L SI+ALVD SS YEYL
Sbjct: 101 MCVC*TDINKDCPRXP--LTSINALVDYSSSYEYL 3
>TC7995 similar to PIR|S63686|S63686 sterol 24-C-methyltransferase -
Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (98%)
Length = 1524
Score = 35.4 bits (80), Expect(2) = 4e-04
Identities = 25/73 (34%), Positives = 32/73 (43%)
Frame = -2
Query: 300 IRAGRLSRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPER 359
+ A RLSR S + PR + PPR P+ P R + P R +TPP +
Sbjct: 914 LAARRLSR----SPESPPRVQDSPPRTFPS*PTRRKRREPPASEPGRFPRTPPPASARGM 747
Query: 360 RRRSRERRRSPDR 372
RS RRR R
Sbjct: 746 SPRSSTRRRMSHR 708
Score = 32.3 bits (72), Expect = 0.64
Identities = 26/100 (26%), Positives = 38/100 (38%)
Frame = +3
Query: 321 SPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRR 380
+PP +R PP P R+S T P +P RRS+ RRSP +
Sbjct: 219 APPNKRANAPPISPADPSPPRKSRTTTTSTGPSSAAP---RRSKPPRRSPISSTPSTISS 389
Query: 381 QHGSNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRP 420
++ V + G + + STR + S RP
Sbjct: 390 PTSTSGVGASPSISPLPSPGNLTATPRASTRSMPSI*SRP 509
Score = 28.9 bits (63), Expect = 7.1
Identities = 24/73 (32%), Positives = 30/73 (40%), Gaps = 14/73 (19%)
Frame = -2
Query: 312 STGALPRPRSPPPRRTPTPPRHRDQTPPK----------RRSAERRDKTPPR-RRSPERR 360
STG P TP+PPR PP+ R R ++PPR + SP R
Sbjct: 1025 STGPTTEAPVVSPDPTPSPPRTPPYAPPQ*CQRSSSALAARRLSRSPESPPRVQDSPPRT 846
Query: 361 RRS---RERRRSP 370
S R +RR P
Sbjct: 845 FPS*PTRRKRREP 807
Score = 26.6 bits (57), Expect(2) = 4e-04
Identities = 22/66 (33%), Positives = 27/66 (40%)
Frame = -3
Query: 388 DVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPPRQKVAITFTEDDYGEDTGE 447
DVGS GG GPT NS ++ G L + VA+ F D GE G
Sbjct: 592 DVGSRVGGDGTHGPT-NSASHIEDLVTRLGLD*IDGMLLVEARGVAVRFPGDGRGEMEGL 416
Query: 448 EDDPIV 453
P+V
Sbjct: 415 APTPLV 398
>TC8296 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;}, partial (55%)
Length = 1276
Score = 42.4 bits (98), Expect = 6e-04
Identities = 49/171 (28%), Positives = 69/171 (39%), Gaps = 11/171 (6%)
Frame = +2
Query: 203 GPQKMKEHKDQPSTRESRRK-GQDERKPKRKKYDSYTPLNSSLSRILREKASTDLRDRPP 261
GP + HK + E R K G R P+R+ D + SS R R ++ R R
Sbjct: 368 GPNAERIHKGRIIESEERAKQGSRSRSPRRRHRDDH---RSSRDRDYRRRS----RSRSY 526
Query: 262 PLLTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGR----------LSRYVAV 311
RGD+ G + D + + L GR SR +V
Sbjct: 527 DKYERGDR-----------HRGRDRDYRRRSRSRSASLDYKGRGRGRYDDEPSRSRSRSV 673
Query: 312 STGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRR 362
+ + P RSP PRR+P+P R PKR ++ R+ PR SP+ R R
Sbjct: 674 DSRS-PARRSPSPRRSPSPKR---SVSPKRSTSPRKS---PRGESPDNRSR 805
>TC14532 similar to UP|Q9FYA8 (Q9FYA8) SC35-like splicing factor SCL30a, 30a
kD, partial (71%)
Length = 1073
Score = 41.6 bits (96), Expect = 0.001
Identities = 31/70 (44%), Positives = 35/70 (49%), Gaps = 8/70 (11%)
Frame = +3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAER-RD--KTPPRRR-----SPERRRRSRERRRS 369
R RSPP R PR+ PP+ RS R RD PP+RR SPE RR SRER +
Sbjct: 465 RRRSPP--RYSRSPRYSRSPPPRHRSRSRSRDYYSPPPKRRDSRSVSPEERRYSRERSYT 638
Query: 370 PDRHGRDEVR 379
P R R
Sbjct: 639 PRSRERSYSR 668
Score = 37.7 bits (86), Expect = 0.015
Identities = 35/116 (30%), Positives = 47/116 (40%), Gaps = 14/116 (12%)
Frame = +3
Query: 325 RRTPTPPRHRDQTP----PKRRSAERRDKTPPRRRSPERRRRSRERRR---SPDRHGRD- 376
R+ PT RHR+++ +RRS R ++P RSP R RSR R R SP RD
Sbjct: 408 RKKPTEMRHRERSRGRSYDRRRSPPRYSRSPRYSRSPPPRHRSRSRSRDYYSPPPKRRDS 587
Query: 377 -----EVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPR-PGSLH 426
E RR S ++ P N + +R A R P +H
Sbjct: 588 RSVSPEERRYSRERSYTPRSRERSYSRSPPYNGGSRSRSRSPAKVAVHSRSPSPIH 755
Score = 35.8 bits (81), Expect = 0.058
Identities = 26/68 (38%), Positives = 30/68 (43%), Gaps = 17/68 (25%)
Frame = +3
Query: 320 RSPPPRRTPT---------PPRHRDQ---TPPKRRSAERRDKTPPRR-----RSPERRRR 362
RSPPPR PP+ RD +P +RR + R TP R RSP
Sbjct: 510 RSPPPRHRSRSRSRDYYSPPPKRRDSRSVSPEERRYSRERSYTPRSRERSYSRSPPYNGG 689
Query: 363 SRERRRSP 370
SR R RSP
Sbjct: 690 SRSRSRSP 713
>TC16930
Length = 520
Score = 38.9 bits (89), Expect = 0.007
Identities = 24/71 (33%), Positives = 38/71 (52%), Gaps = 1/71 (1%)
Frame = +1
Query: 532 FLVVECPTAYNAILGRPSLNTFRAIISTHHLMLKYPWAG-RAVPVRGNLEMARSCYNSSC 590
F++ T+YNAIL PSLN F I T L K+ V ++G+ E+A +CY S
Sbjct: 10 FMLANNLTSYNAILC*PSLNVFDMCILTRRLASKFSTDNTEIVTIQGDQEVAPNCYIMSM 189
Query: 591 RLAREERKRKK 601
+ + ++ K+
Sbjct: 190 WIIPDSQEAKE 222
>TC8088 similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (69%)
Length = 1527
Score = 38.9 bits (89), Expect = 0.007
Identities = 28/79 (35%), Positives = 34/79 (42%), Gaps = 13/79 (16%)
Frame = +3
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKR---RSAERRDKTPPR----------RRSPERRRRS 363
PR P P PPR RDQ PP R +RR PPR R P R RR
Sbjct: 174 PRLLHLQPLPHPHPPRRRDQPPPNHHHPRHRQRRHVLPPRQAPHPLHPKKRPLPPRPRRL 353
Query: 364 RERRRSPDRHGRDEVRRQH 382
R+++P + R+ R H
Sbjct: 354 LWRQKAPPDNQRNHPRLLH 410
>BP054244
Length = 465
Score = 38.9 bits (89), Expect = 0.007
Identities = 25/60 (41%), Positives = 34/60 (56%), Gaps = 1/60 (1%)
Frame = -1
Query: 118 VKQKEKESLKAFLNRFNKEAGDITGLLPDTRLVLATAALAPG-PFLTSLDGKPANTLEEF 176
+KQ +KESL+ FL +F KEA +T L RL L A+ G F TS P+ ++EF
Sbjct: 453 LKQGKKESLRDFLTKFYKEATLVTSLDLKVRLHLLCEAIRMGTSFYTSSAXTPSQDMDEF 274
>TC8148 homologue to PIR|E86216|E86216 protein T23G18.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;}, partial
(75%)
Length = 1527
Score = 38.5 bits (88), Expect = 0.009
Identities = 50/186 (26%), Positives = 73/186 (38%), Gaps = 6/186 (3%)
Frame = +2
Query: 205 QKMKEHKDQPSTRESRRKGQDERKPKRKKYDSYTPLNSSLSRILREKASTDLR-DRPPPL 263
++ H+ +P R+ R RKP+R+ +L R +A +D R D P +
Sbjct: 323 RRQARHRQRPQIRQGRPSAGSYRKPRRRNQSDPAEAADNL----RPRAGSDARLDSPRRV 490
Query: 264 LTRGDKLDSKRFCEFHDSPGHNTDECLNLKDKVEELIRAGRLSRYVAVSTGA-LPRPRSP 322
R ++ R C P ++ V G L R T LPR +P
Sbjct: 491 PGRPRRVF--RGCGGEGLPELRGHPV*PIRGSVPG---GGALDRRPRCPTSRKLPRRLNP 655
Query: 323 PPRRTPTPPRHRDQTPPKRRSAERRDKTPPR-RRSPERRRRSRERRRS---PDRHGRDEV 378
P R P PP P +RRS PPR RSP+ +R+ S P R ++
Sbjct: 656 P*LRPPPPP----PPPHRRRSPTSSPPLPPRPPRSPQGLSHARDFPPSSTLPVRAHVNQT 823
Query: 379 RRQHGS 384
R Q G+
Sbjct: 824 RPQRGA 841
>BP044109
Length = 497
Score = 38.5 bits (88), Expect = 0.009
Identities = 28/73 (38%), Positives = 29/73 (39%), Gaps = 6/73 (8%)
Frame = +3
Query: 316 LPRPRSPPP---RRTPTPPRHRDQTPPKRRSAERRDKTPP---RRRSPERRRRSRERRRS 369
LPR PPP R P PP H Q PP+ R PP R P RR RR
Sbjct: 279 LPRKPPPPPHMARLQPFPPLHPPQLPPRLRPRRALQMAPPSLLRHARPRHRRHLCPRRAG 458
Query: 370 PDRHGRDEVRRQH 382
H R V R H
Sbjct: 459 *QVH-RYAVPRSH 494
Score = 28.9 bits (63), Expect = 7.1
Identities = 12/36 (33%), Positives = 14/36 (38%)
Frame = +2
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPP 352
P P PPP + T P H K + TPP
Sbjct: 110 PPPPPPPPNKNKTTPSHASNATSKSTPRTQTPTTPP 217
>AI967840
Length = 393
Score = 38.1 bits (87), Expect = 0.012
Identities = 22/56 (39%), Positives = 28/56 (49%)
Frame = +2
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDR 372
PR R RR+P P RH + +RR + RR + P RR RR SR R + R
Sbjct: 56 PRARRSQRRRSPPPRRHHRRR--RRRRSRRRTRPPLTRRRRRSRRASRPTRSTSSR 217
Score = 36.6 bits (83), Expect = 0.034
Identities = 21/45 (46%), Positives = 21/45 (46%)
Frame = +2
Query: 336 QTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRR 380
Q P RRS RR P R RRRRSR R R P R RR
Sbjct: 50 QWPRARRSQRRRSPPPRRHHRRRRRRRSRRRTRPPLTRRRRRSRR 184
Score = 31.6 bits (70), Expect = 1.1
Identities = 21/49 (42%), Positives = 26/49 (52%), Gaps = 3/49 (6%)
Frame = +2
Query: 324 PRRTPTPPR-HRDQTPPKRRSAERRDKTPPRRRS--PERRRRSRERRRS 369
P + P R R ++PP RR RR + RRR+ P RRR R RR S
Sbjct: 44 PSQWPRARRSQRRRSPPPRRHHRRRRRRRSRRRTRPPLTRRRRRSRRAS 190
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.325 0.138 0.443
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,210,030
Number of Sequences: 28460
Number of extensions: 536581
Number of successful extensions: 7967
Number of sequences better than 10.0: 367
Number of HSP's better than 10.0 without gapping: 5636
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7193
length of query: 1706
length of database: 4,897,600
effective HSP length: 103
effective length of query: 1603
effective length of database: 1,966,220
effective search space: 3151850660
effective search space used: 3151850660
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0059.7


