

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
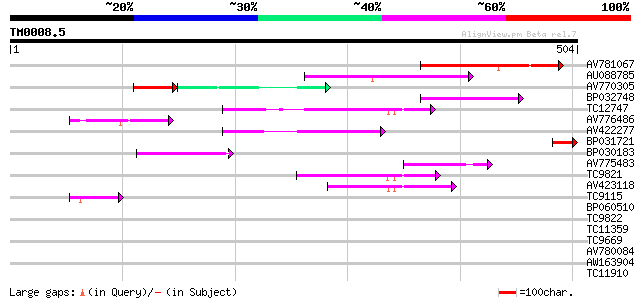
Query= TM0008.5
(504 letters)
Database: LJGI
28,460 sequences; 14,692,800 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AV781067 138 2e-33
AU088785 86 1e-17
AV770305 50 2e-13
BP032748 65 3e-11
TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-l... 61 5e-10
AV776486 59 2e-09
AV422277 54 6e-08
BP031721 52 2e-07
BP030183 52 2e-07
AV775483 49 1e-06
TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 49 3e-06
AV423118 46 1e-05
TC9115 similar to PIR|T00425|T00425 photolyase/blue-light recept... 40 7e-04
BP060510 39 0.002
TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like prot... 39 0.003
TC11359 39 0.003
TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%) 38 0.004
AV780084 37 0.008
AW163904 37 0.008
TC11910 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Saccha... 37 0.010
>AV781067
Length = 538
Score = 138 bits (347), Expect = 2e-33
Identities = 66/129 (51%), Positives = 89/129 (68%), Gaps = 2/129 (1%)
Frame = -2
Query: 366 NGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALAGINYEG 425
NG+RIKTWQGG+GSV + F NIQ+ V+ I IDQ+YC + C N TSAV + ++Y
Sbjct: 525 NGLRIKTWQGGTGSVTNLRFENIQMENVRNCINIDQYYCLTKECRNQTSAVHVNDVSYIN 346
Query: 426 IKGTYTVK--PVHFACSDSLPCVDVSLTSVELKPVQEQYHLYEPFCWQTYGQLMTPTTPP 483
IKGTY V+ P+HFACSDS+ C +++L+ VEL P + + L + FCW YG T T PP
Sbjct: 345 IKGTYDVRTPPIHFACSDSVACTNITLSDVELLPYEGEL-LEDSFCWNAYGTQETITVPP 169
Query: 484 IDCIQIGKP 492
IDC++ G+P
Sbjct: 168 IDCLREGEP 142
>AU088785
Length = 773
Score = 85.9 bits (211), Expect = 1e-17
Identities = 58/175 (33%), Positives = 80/175 (45%), Gaps = 25/175 (14%)
Frame = +2
Query: 263 PTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNS- 321
P++L F + V I N H+ C V + + + +P SPNTDGIH+ +S
Sbjct: 14 PSSLFFMDXSNVVVQNIKSLNPKGFHMFVTKCTNVRLRKLKLIAPETSPNTDGIHISSSI 193
Query: 322 ----------------------KDVLIHSSKMACGHGISIGSLGKDNTRACVSNITIRDV 359
++V I+ K GHGISIGSLGK V I I +
Sbjct: 194 NVIIARNTIQTGDDCISMIXGSENVFINRLKCGPGHGISIGSLGKYAEXREVKGIXIXNS 373
Query: 360 NMHNTMNGVRIKTWQGG-SGSVQGVLFSNIQVSEVQLPIVIDQFY-CDKRSCTNH 412
+ T NG+RIK+W G+ + FSNI + V+ PI+IDQ Y C NH
Sbjct: 374 ALIGTTNGLRIKSWPDKYGGAASXISFSNITMENVKNPIIIDQXYQCSPNCQKNH 538
>AV770305
Length = 442
Score = 50.4 bits (119), Expect(2) = 2e-13
Identities = 37/137 (27%), Positives = 54/137 (39%), Gaps = 1/137 (0%)
Frame = +2
Query: 150 VPAAYVFYVGPIS-FSGPYCKPNIVFQLDGTIIAPTNSNAWGGGLLQWLEFSKLVGITIQ 208
+P F V I+ GP +I QL G I+AP+ +AWG +W+ + +T+
Sbjct: 134 IPGGKTFLVTSITTLPGPCKSSSINIQLQGNIVAPS-FDAWGADKSKWISIHSVNHLTMN 310
Query: 209 GKGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSLPVQNEMGRQMPAVKPTALRF 268
G G IDG GS W+ R +PT+L F
Sbjct: 311 GGGKIDGLGSSGWE---------------------------------RCRTCARPTSLSF 391
Query: 269 YGSFSPTVTGITIQNSP 285
Y TV+ + + NSP
Sbjct: 392 YSGNGLTVSNLHLANSP 442
Score = 42.0 bits (97), Expect(2) = 2e-13
Identities = 18/40 (45%), Positives = 30/40 (75%), Gaps = 1/40 (2%)
Frame = +1
Query: 111 STMFNVLDFGAKGDGRTDDTKAFQATCAMACK-VEASTMV 149
+T++NV+ +GAKGDG++DD+ AF + +AC+ E T+V
Sbjct: 13 NTIYNVMHYGAKGDGKSDDSHAFVSAWKVACESAEVGTLV 132
>BP032748
Length = 453
Score = 64.7 bits (156), Expect = 3e-11
Identities = 35/93 (37%), Positives = 52/93 (55%), Gaps = 2/93 (2%)
Frame = -2
Query: 366 NGVRIKTWQGG-SGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTNHTSAVALAGINYE 424
NG+RIK+W G+ + FSNI + V+ PI+IDQ Y +C S V + I++
Sbjct: 452 NGLRIKSWPDKYGGAASEISFSNITMENVKNPIIIDQEYQCSPNCQKKPSLVRIRDIHFA 273
Query: 425 GIKGTYTVK-PVHFACSDSLPCVDVSLTSVELK 456
+KGT T V F CS PC+ ++L ++LK
Sbjct: 272 NVKGTTTSPIAVDFRCSKLYPCMGITLRDIDLK 174
>TC12747 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-like
protein, partial (43%)
Length = 624
Score = 60.8 bits (146), Expect = 5e-10
Identities = 53/223 (23%), Positives = 91/223 (40%), Gaps = 34/223 (15%)
Frame = +2
Query: 190 GGGLLQWLEFSKLVGITIQG-KGVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSL 248
GG + + L + I G G IDG+GSVWW D LN++
Sbjct: 44 GGRYRSLIYGNNLTDVVITGYDGTIDGQGSVWWDLISTGD-----------LNYS----- 175
Query: 249 PVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPG 308
+P + GS + T++ +TI NSP + C V +H++++ +P
Sbjct: 176 -------------RPHIIELIGSDNITISNLTILNSPSWGIHPVYCRYVQIHNITVHAPP 316
Query: 309 DSPNTDGIHLQNSKDVLIHSSKMACGH------------GISIG---------------- 340
+SP+T GI +S+ + I S ++ GH G++ G
Sbjct: 317 ESPHTSGIVPDSSEYICIDKSNISTGHDAIALKSGWDEYGVAYGKPTLNVHISSVYLQSS 496
Query: 341 -----SLGKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSG 378
+ G + + +SNI V++ N+ G+ +KT +G G
Sbjct: 497 SGAGLAFGSEMSGG-ISNIIAEQVHIINSKIGIELKTTKGRGG 622
>AV776486
Length = 409
Score = 58.9 bits (141), Expect = 2e-09
Identities = 35/94 (37%), Positives = 43/94 (45%), Gaps = 2/94 (2%)
Frame = +1
Query: 54 NSHGNSHNQHGGGGSSSHSKPKPPPPTHKKTPSSPLFPKPKVDP--PSTPPSNDYNGGSS 111
N G +H + P P P H P+S P P PS P ND G S
Sbjct: 142 NVEGRNHYHK----KQNKKSPAPKSPAHSPAPASDTSPPSSTSPSVPSDPYPND-PGDSG 306
Query: 112 TMFNVLDFGAKGDGRTDDTKAFQATCAMACKVEA 145
+F+V+ FGA GDG DDT AF+ AC VE+
Sbjct: 307 CVFDVMSFGAVGDGSADDTAAFREAWKAACAVES 408
>AV422277
Length = 454
Score = 53.9 bits (128), Expect = 6e-08
Identities = 36/146 (24%), Positives = 64/146 (43%), Gaps = 1/146 (0%)
Frame = +2
Query: 190 GGGLLQWLEFSKLVGITIQGK-GVIDGRGSVWWQDFPYDDPIDDEEKLIVPLNHTQKPSL 248
GG + ++ + + I G+ G IDG+G VWW +
Sbjct: 77 GGRYVSFIHGDGVRDVVITGENGTIDGQGDVWWNMWRQ---------------------- 190
Query: 249 PVQNEMGRQMPAVKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPG 308
R + +P + F S + ++ + ++SP ++ C+ V+V V+I +P
Sbjct: 191 -------RTLQFTRPNLVEFVNSRNIIISNVIFKDSPFWNIHPVYCSNVVVRYVTILAPR 349
Query: 309 DSPNTDGIHLQNSKDVLIHSSKMACG 334
DSPNTDG+ +S +V I S ++ G
Sbjct: 350 DSPNTDGVDPDSSSNVCIEDSYISTG 427
>BP031721
Length = 486
Score = 52.0 bits (123), Expect = 2e-07
Identities = 22/22 (100%), Positives = 22/22 (100%)
Frame = -2
Query: 483 PIDCIQIGKPSSNRIQTNHDIC 504
PIDCIQIGKPSSNRIQTNHDIC
Sbjct: 485 PIDCIQIGKPSSNRIQTNHDIC 420
>BP030183
Length = 483
Score = 52.0 bits (123), Expect = 2e-07
Identities = 29/88 (32%), Positives = 45/88 (50%), Gaps = 1/88 (1%)
Frame = +3
Query: 113 MFNVLDFGAKGDGRTDDTKAFQATCAMACKVEASTMVVPAAYVFYVGPISFSGPYCKPN- 171
++NV+ FGAK DG D T+AF C A ++ A F V + F GP P+
Sbjct: 222 IYNVMSFGAKPDGVFDSTQAFMTAWQTVCHSPAPGRILVPAGRFLVSSMFFQGPCLSPHP 401
Query: 172 IVFQLDGTIIAPTNSNAWGGGLLQWLEF 199
+ Q++GT++A + + + G WL F
Sbjct: 402 VTLQVEGTVLASADLSEFENG--DWLMF 479
>AV775483
Length = 503
Score = 49.3 bits (116), Expect = 1e-06
Identities = 25/79 (31%), Positives = 42/79 (52%)
Frame = -2
Query: 351 VSNITIRDVNMHNTMNGVRIKTWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCT 410
V + + D + T N RIKT GGSG + + F I +++ PI+ID +Y +
Sbjct: 484 VEEVYVYDCSFTGTQNPARIKTLSGGSGYARKITFEKITLTKAGDPIIIDPYYAPE---- 317
Query: 411 NHTSAVALAGINYEGIKGT 429
S+V ++ + + GI+GT
Sbjct: 316 -DDSSVPVSDVTFRGIRGT 263
>TC9821 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (61%)
Length = 1080
Score = 48.5 bits (114), Expect = 3e-06
Identities = 38/161 (23%), Positives = 70/161 (42%), Gaps = 33/161 (20%)
Frame = +3
Query: 256 RQMPAVKPTALRFYGSFSPTVTGITIQNSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDG 315
+Q+ +P + S + ++ +T+ NSP ++ C+ V+V ++I +P SPNTDG
Sbjct: 45 KQLQYTRPYLIEIMYSDNVQISNLTLVNSPSWNIHPVYCSNVIVQGITILAPVTSPNTDG 224
Query: 316 IHLQNSKDVLIHSSKMACG------------HGISIG---------------------SL 342
I+ + + I + G +GIS G +L
Sbjct: 225 INPDSCTNTRIEDCYIVSGDDCVAVKSGWDEYGISYGMPTKHLVIRRLTCISPTSAVIAL 404
Query: 343 GKDNTRACVSNITIRDVNMHNTMNGVRIKTWQGGSGSVQGV 383
G + + + ++ D+ N+ +GVRIKT G G V+ +
Sbjct: 405 GSEMSGG-IEDVRAEDILAINSESGVRIKTAVGRGGYVRDI 524
>AV423118
Length = 472
Score = 46.2 bits (108), Expect = 1e-05
Identities = 32/148 (21%), Positives = 67/148 (44%), Gaps = 33/148 (22%)
Frame = +2
Query: 283 NSPQCHLKFDNCNGVLVHDVSISSPGDSPNTDGIHLQNSKDVLIHSSKMACGH------- 335
N+P + C+ V +H+++I +P +SP+T G+ +S V I +A G+
Sbjct: 5 NAPAYSIHPVYCSYVHIHNLTIFAPPESPDTVGLVPDSSDHVCIEDCVIATGYDAIALKS 184
Query: 336 -----GISIG---------------------SLGKDNTRACVSNITIRDVNMHNTMNGVR 369
GI+ G + G D + +SN+ + + ++HN+ G+
Sbjct: 185 GWDEYGIAYGRPTENVHIRRVDLQASSGSALAFGSDMSGG-ISNVLVENAHLHNSNGGIE 361
Query: 370 IKTWQGGSGSVQGVLFSNIQVSEVQLPI 397
+T +G G ++ ++ S++++ V I
Sbjct: 362 FRTTRGRGGYMKDIIISDVEMKNVYTAI 445
>TC9115 similar to PIR|T00425|T00425 photolyase/blue-light receptor (PHR2)
[imported] - Arabidopsis thaliana
{Arabidopsis thaliana;}, partial (55%)
Length = 1018
Score = 40.4 bits (93), Expect = 7e-04
Identities = 23/52 (44%), Positives = 29/52 (55%), Gaps = 4/52 (7%)
Frame = +1
Query: 54 NSHGNSHN--QHGGGGSSSHSKPKPPP-PT-HKKTPSSPLFPKPKVDPPSTP 101
+S+ NSH+ H + S S+PKPPP PT H P P P P +PPS P
Sbjct: 196 SSNPNSHHLLPHNNNPTRSKSQPKPPPSPTSHSPPPPHPRRPNPPSNPPSPP 351
Score = 27.3 bits (59), Expect = 6.0
Identities = 16/41 (39%), Positives = 21/41 (51%), Gaps = 2/41 (4%)
Frame = +1
Query: 74 PKPPPPTHKKTPSSPLFPKPKVDPP--STPPSNDYNGGSST 112
P PP P+ TP +P P PP + PPS+ GS+T
Sbjct: 340 PSPPTPSAPLTP*APTAPSTLPTPPHSAAPPSS----GSAT 450
Score = 26.9 bits (58), Expect = 7.8
Identities = 14/35 (40%), Positives = 16/35 (45%), Gaps = 5/35 (14%)
Frame = -2
Query: 75 KPPPPTHKKTPS-----SPLFPKPKVDPPSTPPSN 104
K PPT K P SP P PPS+PP +
Sbjct: 870 KAYPPTEKANPQHGTKCSPKSTSPPHSPPSSPPQS 766
>BP060510
Length = 462
Score = 38.9 bits (89), Expect = 0.002
Identities = 26/59 (44%), Positives = 31/59 (52%), Gaps = 2/59 (3%)
Frame = +1
Query: 78 PPTHKKTPS-SPLFPKPKVDP-PSTPPSNDYNGGSSTMFNVLDFGAKGDGRTDDTKAFQ 134
PP PS SP FP DP P+ P + NG +F+V GA GDG DDT AF+
Sbjct: 307 PPNDPSPPSISPSFPS---DPYPNDPQDSSSNG----IFDVRSXGAVGDGSADDTPAFR 462
>TC9822 similar to UP|Q84LI7 (Q84LI7) Polygalacturonase-like protein,
partial (40%)
Length = 915
Score = 38.5 bits (88), Expect = 0.003
Identities = 31/130 (23%), Positives = 52/130 (39%), Gaps = 20/130 (15%)
Frame = +1
Query: 115 NVLDFGAKGDGRTDDTKAFQATCAMACKVEA---STMVVPAAYVFYVGPISFSGPYCKPN 171
+V DFG GDG T +T+AF+ + + S + VP + G + + +
Sbjct: 160 SVTDFGGVGDGTTLNTQAFRKAIEHLSQYSSNGGSQLYVPPGR-WLTGSFNLTSHF---T 327
Query: 172 IVFQLDGTIIAPTNSNAW----------------GGGLLQWLEFSKLVGITIQG-KGVID 214
+ D I+ + N W GG + + L + I G G +D
Sbjct: 328 LFIHKDAVILGSQDENEWPVIDPLPSYGRGRDTQGGRFSSLICGTHLTDVIITGDNGTLD 507
Query: 215 GRGSVWWQDF 224
G+G +WW+ F
Sbjct: 508 GQGDLWWKKF 537
>TC11359
Length = 544
Score = 38.5 bits (88), Expect = 0.003
Identities = 31/141 (21%), Positives = 56/141 (38%), Gaps = 25/141 (17%)
Frame = -3
Query: 372 TWQGGSGSVQGVLFSNIQVSEVQLPIVIDQFYCDKRSCTN----------------HTSA 415
T +GG G + +++ NI V++ P+ IDQ Y N H S
Sbjct: 443 TMKGGKGYAKAIIYENIIVNQTNYPVYIDQHYMRTPEKVNILSYCLLFVDLFGDNLHHSF 264
Query: 416 VALAGINYEGIKGT-YTVKPVHFACSDS----LPCVDVSLTSVELKPVQ-EQYHLYEP-- 467
L + E +K T T + +H C++ L C + +++L+ + L +P
Sbjct: 263 PLLFWLQKEALKVTDVTFRNIHGTCTNEHAVVLDCAKIGCDNIKLQQINITSIDLDKPAS 84
Query: 468 -FCWQTYGQLMTPTTPPIDCI 487
C +G +P + C+
Sbjct: 83 AICHNVHGTATDVISPRVTCL 21
>TC9669 similar to UP|Q9LQA7 (Q9LQA7) F4N2.10, partial (16%)
Length = 472
Score = 37.7 bits (86), Expect = 0.004
Identities = 23/61 (37%), Positives = 28/61 (45%), Gaps = 12/61 (19%)
Frame = +3
Query: 53 GNSHGNSHNQHGGGGS--SSHSKPKPP--------PPTHKKTPSSPLFPKPKVDP--PST 100
G HG++ HGGGG + S P PP PP TP++P P P PST
Sbjct: 273 GGGHGSTPPSHGGGGGYYNPPSSPSPPKGGNCGSSPPHDPSTPTTPSTPHTPSTPHTPST 452
Query: 101 P 101
P
Sbjct: 453 P 455
>AV780084
Length = 527
Score = 37.0 bits (84), Expect = 0.008
Identities = 22/63 (34%), Positives = 35/63 (54%), Gaps = 1/63 (1%)
Frame = -3
Query: 413 TSAVALAGINYEGIKGTY-TVKPVHFACSDSLPCVDVSLTSVELKPVQEQYHLYEPFCWQ 471
TSAV + I++ I+GT T + + FACSD+ PC + L ++ L+ +CWQ
Sbjct: 516 TSAVQVRNISFTDIQGTSATEEAIKFACSDASPCEGLYL*NIFLESCFGGN--TRSYCWQ 343
Query: 472 TYG 474
+G
Sbjct: 342 AHG 334
>AW163904
Length = 416
Score = 37.0 bits (84), Expect = 0.008
Identities = 17/31 (54%), Positives = 18/31 (57%)
Frame = +2
Query: 49 FKKKGNSHGNSHNQHGGGGSSSHSKPKPPPP 79
FKK S+ NS N GGGG SK PPPP
Sbjct: 20 FKKSKFSNQNSSNPGGGGGGGDSSKEAPPPP 112
Score = 28.9 bits (63), Expect = 2.1
Identities = 20/62 (32%), Positives = 29/62 (46%), Gaps = 4/62 (6%)
Frame = -3
Query: 22 IWSSNFETCIARRGKHWRQSRTVSASLFKKKGN----SHGNSHNQHGGGGSSSHSKPKPP 77
I SN T I + W R+ S+S+ + S ++ + GGGG+S P PP
Sbjct: 249 IMRSNEFTTICICSQ*WLYLRSTSSSMARSLTFLL*CSILSAADNGGGGGASFEESPPPP 70
Query: 78 PP 79
PP
Sbjct: 69 PP 64
>TC11910 similar to GB|CAA54498.1|602891|SCIILDNA YBL0422 {Saccharomyces
cerevisiae;} , partial (6%)
Length = 638
Score = 36.6 bits (83), Expect = 0.010
Identities = 33/109 (30%), Positives = 46/109 (41%), Gaps = 7/109 (6%)
Frame = +1
Query: 45 SASLFKKKGNSHGNSHNQHGGGGSSSHS--KPKPPPPTHKKTPSSPLFPKPKVDPPSTPP 102
S+SL +S +S + H SSHS +P PPPP+ PS P FP P PP
Sbjct: 28 SSSLPPSSSSSSSSSTSLHLSP-CSSHSLPQPSPPPPSKSHFPSLPRFPSKCRHKPPKPP 204
Query: 103 SNDYNGGSSTMFNVLD-----FGAKGDGRTDDTKAFQATCAMACKVEAS 146
+F ++D G G G DT+ AT ++ + S
Sbjct: 205 ---------RLFMIIDPILLLNGFGGAGFYLDTQTLLATVSVLAAIALS 324
Database: LJGI
Posted date: Jul 30, 2004 11:16 AM
Number of letters in database: 14,692,800
Number of sequences in database: 28,460
Lambda K H
0.318 0.135 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,908,036
Number of Sequences: 28460
Number of extensions: 210870
Number of successful extensions: 4570
Number of sequences better than 10.0: 389
Number of HSP's better than 10.0 without gapping: 2868
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3999
length of query: 504
length of database: 4,897,600
effective HSP length: 94
effective length of query: 410
effective length of database: 2,222,360
effective search space: 911167600
effective search space used: 911167600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Lotus: description of TM0008.5


