

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0400a.3
(419 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
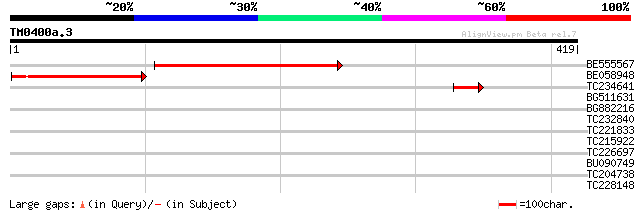
BE555567 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsi... 268 4e-72
BE058948 165 3e-41
TC234641 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2,... 43 2e-04
BG511631 similar to GP|18087557|gb| AT3g27930/K24A2_2 {Arabidops... 34 0.11
BG882216 33 0.24
TC232840 32 0.70
TC221833 UP|Q9LWW4 (Q9LWW4) ESTs C26056(C11550), partial (10%) 30 2.6
TC215922 29 3.5
TC226697 weakly similar to UP|Q7RHM7 (Q7RHM7) Cyclophilin-RNA in... 29 4.5
BU090749 28 5.9
TC204738 28 5.9
TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein ... 28 5.9
>BE555567 similar to GP|18087557|gb AT3g27930/K24A2_2 {Arabidopsis thaliana},
partial (28%)
Length = 422
Score = 268 bits (684), Expect = 4e-72
Identities = 127/139 (91%), Positives = 134/139 (96%)
Frame = +2
Query: 108 SCAYYPRYGFGAFGVFPLLLRKREFSEDYGLMGLRYGSGNLSFGVTLMPFAKKDELPKSA 167
+CAYYP+YGFGAFGVFPLLL+KRE S+DYGLMGLRYGSGNLSFGVTL+PFA KDELPKSA
Sbjct: 5 ACAYYPKYGFGAFGVFPLLLKKRESSQDYGLMGLRYGSGNLSFGVTLLPFAMKDELPKSA 184
Query: 168 WLVSKMGRLTAGVQYEPQHGNAKLSNLMNWSCAMAYGVGSGSPLSPSFNFSLELVKSSQF 227
WLVSKMGRLTAGVQYEPQ GNAKLSNLMNWSCAM YGVGSGSPL PSFNF+LELVKSSQF
Sbjct: 185 WLVSKMGRLTAGVQYEPQQGNAKLSNLMNWSCAMGYGVGSGSPLCPSFNFNLELVKSSQF 364
Query: 228 VASFYQHMVVQRRVKNPLE 246
+ASFYQHMVVQRRVKNPLE
Sbjct: 365 IASFYQHMVVQRRVKNPLE 421
>BE058948
Length = 411
Score = 165 bits (418), Expect = 3e-41
Identities = 79/100 (79%), Positives = 87/100 (87%)
Frame = +3
Query: 2 FRKEVKPPPPPLVVLVPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFNDYFQSPKHFT 61
F +E PPPPP VVLVPPLFDFPPLAARNRML SSYDVVFGKLAL LF DYF ++FT
Sbjct: 57 FSREPAPPPPP-VVLVPPLFDFPPLAARNRMLHSSYDVVFGKLALTSLFEDYFHQARNFT 233
Query: 62 TRIMFKPIDEPHVDLIATVSGPLDHKPEESITGNALFRWQ 101
TRIM KPI++PHVDLIATVSGPLDHKP++SI G+ALFRWQ
Sbjct: 234 TRIMLKPIEDPHVDLIATVSGPLDHKPKDSIAGDALFRWQ 353
>TC234641 weakly similar to UP|Q8W106 (Q8W106) AT3g27930/K24A2_2, partial
(16%)
Length = 384
Score = 43.1 bits (100), Expect = 2e-04
Identities = 19/22 (86%), Positives = 22/22 (99%)
Frame = +3
Query: 329 ADGQIQYGFGIQSESLREASFQ 350
ADG++QYGFGIQSESLREAS+Q
Sbjct: 12 ADGKMQYGFGIQSESLREASYQ 77
>BG511631 similar to GP|18087557|gb| AT3g27930/K24A2_2 {Arabidopsis
thaliana}, partial (3%)
Length = 297
Score = 34.3 bits (77), Expect = 0.11
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = -2
Query: 214 SFNFSLELVKSSQFVASFYQHMVVQRRV 241
S + + + QF+ASFYQHM VQRRV
Sbjct: 152 SILLKFQSLNNLQFIASFYQHMAVQRRV 69
>BG882216
Length = 398
Score = 33.1 bits (74), Expect = 0.24
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = +2
Query: 370 VVSYNILGVDSASKHQDLYSNIRHSFQN 397
V SYNILG +AS+H DLY N+ + N
Sbjct: 302 VASYNILGGRNASQHSDLYVNVPSRYIN 385
>TC232840
Length = 449
Score = 31.6 bits (70), Expect = 0.70
Identities = 12/22 (54%), Positives = 15/22 (67%)
Frame = +1
Query: 8 PPPPPLVVLVPPLFDFPPLAAR 29
P PPP++ L+PP F F PL R
Sbjct: 76 PLPPPMLHLLPPFFSFSPLCIR 141
>TC221833 UP|Q9LWW4 (Q9LWW4) ESTs C26056(C11550), partial (10%)
Length = 542
Score = 29.6 bits (65), Expect = 2.6
Identities = 14/43 (32%), Positives = 22/43 (50%), Gaps = 6/43 (13%)
Frame = -3
Query: 8 PPPPPLVV------LVPPLFDFPPLAARNRMLESSYDVVFGKL 44
PPPPP V+ L+ P+ PP A N+ +E+ + + L
Sbjct: 186 PPPPPFVLLLVARTLIDPVLSTPPCFALNKKMETELEPLLESL 58
>TC215922
Length = 802
Score = 29.3 bits (64), Expect = 3.5
Identities = 12/20 (60%), Positives = 14/20 (70%)
Frame = +1
Query: 6 VKPPPPPLVVLVPPLFDFPP 25
+ PP PP VL+PPLF F P
Sbjct: 139 LSPPFPPQSVLLPPLFSFLP 198
>TC226697 weakly similar to UP|Q7RHM7 (Q7RHM7) Cyclophilin-RNA interacting
protein, partial (11%)
Length = 488
Score = 28.9 bits (63), Expect = 4.5
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 2/51 (3%)
Frame = +2
Query: 3 RKEVKPPPPPLVVL--VPPLFDFPPLAARNRMLESSYDVVFGKLALRCLFN 51
R+ + PP PP L PP+F PPL + + +L S V L L N
Sbjct: 152 RRHLAPPLPPPSSLRPSPPVFSPPPLLSSSDLLSGSEHVALRPLDFHFLSN 304
>BU090749
Length = 405
Score = 28.5 bits (62), Expect = 5.9
Identities = 11/46 (23%), Positives = 24/46 (51%)
Frame = -2
Query: 312 SWWKPSFTFSISATRDRADGQIQYGFGIQSESLREASFQSIFTILN 357
+WW + + + A+G++ + G+QS SL E F++++
Sbjct: 404 AWWLEFSEVASALGEEEAEGEVDHKTGLQSSSLMEERIWKAFSMVS 267
>TC204738
Length = 940
Score = 28.5 bits (62), Expect = 5.9
Identities = 18/56 (32%), Positives = 26/56 (46%)
Frame = -3
Query: 86 HKPEESITGNALFRWQRILRMRSCAYYPRYGFGAFGVFPLLLRKREFSEDYGLMGL 141
H+P E + N F W+R R C + R G + P L + FS GL+G+
Sbjct: 260 HRPNELVAPNKGFLWRRWRSSRVCIHRRRRGEPVRFLLPPKLLQLRFS---GLVGI 102
>TC228148 similar to UP|Q7YXC8 (Q7YXC8) C. elegans GRD-1 protein
(Corresponding sequence R08B4.1b), partial (3%)
Length = 1180
Score = 28.5 bits (62), Expect = 5.9
Identities = 19/52 (36%), Positives = 25/52 (47%), Gaps = 9/52 (17%)
Frame = -1
Query: 8 PPPPPLVVLV---------PPLFDFPPLAARNRMLESSYDVVFGKLALRCLF 50
PPPPPL++L+ PPL PP A +L ++F K A LF
Sbjct: 568 PPPPPLLLLLLPPPPPLPPPPLPPLPPFDASAEVLRL---ILFKKSAAAFLF 422
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,813,624
Number of Sequences: 63676
Number of extensions: 274270
Number of successful extensions: 2447
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 2085
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2388
length of query: 419
length of database: 12,639,632
effective HSP length: 100
effective length of query: 319
effective length of database: 6,272,032
effective search space: 2000778208
effective search space used: 2000778208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0400a.3


