

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0249.1
(68 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
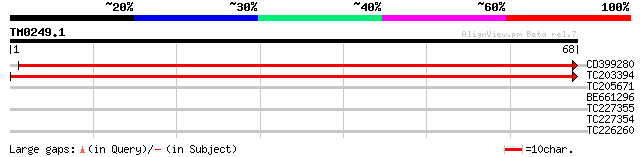
CD399280 weakly similar to GP|16648801|gb At2g24280/F27D4.19 {Ar... 116 2e-27
TC203394 weakly similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X ca... 91 1e-19
TC205671 similar to GB|AAO23606.1|27764970|BT003041 At2g27920/T1... 26 4.5
BE661296 25 5.9
TC227355 similar to UP|TSSP_HUMAN (Q9NQE7) Thymus-specific serin... 25 5.9
TC227354 25 7.7
TC226260 similar to UP|O22285 (O22285) Expressed protein (At2g39... 25 7.7
>CD399280 weakly similar to GP|16648801|gb At2g24280/F27D4.19 {Arabidopsis
thaliana}, partial (20%)
Length = 482
Score = 116 bits (291), Expect = 2e-27
Identities = 55/67 (82%), Positives = 62/67 (92%)
Frame = -1
Query: 2 IEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKWL 61
IE+ LKRSASNIIF+NGLRDPWS GGVLK ISKT+VAIVAK+GAHH DLRYS+KEDP+WL
Sbjct: 383 IERVLKRSASNIIFFNGLRDPWSAGGVLKTISKTIVAIVAKKGAHHVDLRYSSKEDPQWL 204
Query: 62 KDVRIKE 68
KDVR +E
Sbjct: 203 KDVRTQE 183
>TC203394 weakly similar to UP|Q9FLH1 (Q9FLH1) Lysosomal Pro-X
carboxypeptidase, partial (50%)
Length = 1017
Score = 90.5 bits (223), Expect = 1e-19
Identities = 43/68 (63%), Positives = 52/68 (76%)
Frame = +1
Query: 1 DIEKDLKRSASNIIFYNGLRDPWSGGGVLKNISKTLVAIVAKEGAHHRDLRYSTKEDPKW 60
DI LK+ SNIIF NGL DPWSGGGVL+NIS+++V++V +EGAHH DLR STK DP W
Sbjct: 562 DIHATLKKFGSNIIFSNGLLDPWSGGGVLQNISESVVSLVTEEGAHHIDLRSSTKNDPDW 741
Query: 61 LKDVRIKE 68
L + R E
Sbjct: 742 LVEQRETE 765
>TC205671 similar to GB|AAO23606.1|27764970|BT003041 At2g27920/T1E2.16
{Arabidopsis thaliana;} , partial (75%)
Length = 1695
Score = 25.8 bits (55), Expect = 4.5
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +2
Query: 1 DIEKDLKRSASNIIFYNGLRDPWSGGGVL 29
D+E ++ S++N+ FYN L+D S L
Sbjct: 752 DLENEIVASSNNVDFYNFLQDSKSDSDTL 838
>BE661296
Length = 308
Score = 25.4 bits (54), Expect = 5.9
Identities = 11/22 (50%), Positives = 15/22 (68%)
Frame = -1
Query: 33 SKTLVAIVAKEGAHHRDLRYST 54
+KTL+ VAK+G H LR +T
Sbjct: 242 NKTLILYVAKDGQGHHPLRSNT 177
>TC227355 similar to UP|TSSP_HUMAN (Q9NQE7) Thymus-specific serine protease
precursor , partial (5%)
Length = 1722
Score = 25.4 bits (54), Expect = 5.9
Identities = 10/17 (58%), Positives = 12/17 (69%)
Frame = +2
Query: 7 KRSASNIIFYNGLRDPW 23
K + S IIF NG +DPW
Sbjct: 1247 KIAGSKIIFTNGSQDPW 1297
>TC227354
Length = 698
Score = 25.0 bits (53), Expect = 7.7
Identities = 9/17 (52%), Positives = 12/17 (69%)
Frame = +1
Query: 7 KRSASNIIFYNGLRDPW 23
K + S I+F NG +DPW
Sbjct: 250 KIAGSKIVFANGSQDPW 300
>TC226260 similar to UP|O22285 (O22285) Expressed protein (At2g39750/T5I7.5),
partial (79%)
Length = 2458
Score = 25.0 bits (53), Expect = 7.7
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = -3
Query: 34 KTLVAIVAKEGAHHRDLRYSTKEDPK 59
KT + EGA+H+ Y+ EDP+
Sbjct: 1847 KTSERVCINEGAYHKPRPYNLHEDPQ 1770
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.136 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,954,849
Number of Sequences: 63676
Number of extensions: 25757
Number of successful extensions: 109
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 109
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 109
length of query: 68
length of database: 12,639,632
effective HSP length: 44
effective length of query: 24
effective length of database: 9,837,888
effective search space: 236109312
effective search space used: 236109312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 52 (24.6 bits)
Lotus: description of TM0249.1


