

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
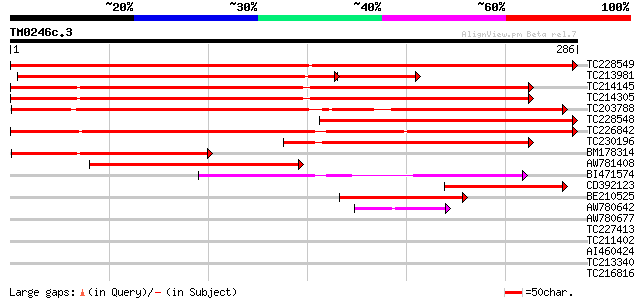
Query= TM0246c.3
(286 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC228549 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {A... 514 e-146
TC213981 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {A... 292 6e-91
TC214145 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated pro... 270 4e-73
TC214305 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associa... 269 1e-72
TC203788 similar to GB|AAL91270.1|19699320|AY090367 AT3g12090/T2... 258 2e-69
TC228548 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {A... 234 3e-62
TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associa... 233 1e-61
TC230196 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associa... 132 1e-31
BM178314 weakly similar to PIR|T02906|T029 senescence-associated... 111 3e-25
AW781408 similar to GP|19699320|gb| AT3g12090/T21B14_110 {Arabid... 103 1e-22
BI471574 100 6e-22
CD392123 71 5e-13
BE210525 70 1e-12
AW780642 55 5e-08
AW780677 33 0.11
TC227413 similar to UP|Q7PS25 (Q7PS25) ENSANGP00000020753 (Fragm... 29 2.1
TC211402 28 3.7
AI460424 28 4.8
TC213340 28 4.8
TC216816 weakly similar to UP|KHLX_HUMAN (Q9Y2M5) Kelch-like pro... 28 4.8
>TC228549 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;} , complete
Length = 1010
Score = 514 bits (1323), Expect = e-146
Identities = 247/286 (86%), Positives = 264/286 (91%)
Frame = +2
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
MRTSNHLIG+LNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG
Sbjct: 11 MRTSNHLIGLLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 190
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRV+NR Y++YYL+DYSG
Sbjct: 191 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVMNRAYLEYYLEDYSG 370
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WLEERVAS SYWGKIASC+RDSKVC +MGR NG+P+ D+FYL LT +QSGCCKPPT
Sbjct: 371 WLEERVASESYWGKIASCIRDSKVCGRMGRT-VNGMPQTPDMFYLTHLTPIQSGCCKPPT 547
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
DCGY+YQNETVW GSGLM N DC++WSNDQ LCY CDSCKAGVLASLKKSWRKVSVI
Sbjct: 548 DCGYVYQNETVWIPGSGLMGTNADCTRWSNDQEQLCYACDSCKAGVLASLKKSWRKVSVI 727
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHPSHFQL 286
NIVVMIILVIVYIIAYAAYRNN++MDNDEPYGEARMTK+ PS F L
Sbjct: 728 NIVVMIILVIVYIIAYAAYRNNRKMDNDEPYGEARMTKAQPSAFHL 865
>TC213981 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;} , partial (62%)
Length = 625
Score = 292 bits (747), Expect(2) = 6e-91
Identities = 141/162 (87%), Positives = 152/162 (93%)
Frame = +1
Query: 5 NHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGA 64
NHLIG+LNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGA
Sbjct: 1 NHLIGLLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGFAGA 180
Query: 65 CYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGWLEE 124
CYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRV+NR Y++YYL+DYSGWLEE
Sbjct: 181 CYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVMNRAYLEYYLEDYSGWLEE 360
Query: 125 RVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLR 166
RVAS SYWGKI SCVRDSK C +MG + NG+PE +D+FY +
Sbjct: 361 RVASDSYWGKIVSCVRDSKACGRMG-ITINGMPETSDMFYXK 483
Score = 60.1 bits (144), Expect(2) = 6e-91
Identities = 28/43 (65%), Positives = 31/43 (71%), Gaps = 1/43 (2%)
Frame = +2
Query: 166 RKLTSVQSGCCKPPTDCGYIYQNET-VWNLGSGLMSANPDCSK 207
R LT +QS CCKPPTDCGY+YQNET V G M ANPD +K
Sbjct: 482 RHLTPIQSXCCKPPTDCGYVYQNETSVDPQDQGXMGANPDWTK 610
>TC214145 similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated protein-like,
partial (87%)
Length = 1867
Score = 270 bits (691), Expect = 4e-73
Identities = 129/265 (48%), Positives = 185/265 (69%), Gaps = 1/265 (0%)
Frame = +1
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN++IG+LNFLTFLLSIPIL G+WLS + T+C ++L+ P+I +GV +M+VSLAG
Sbjct: 862 VRFSNNVIGLLNFLTFLLSIPILVAGVWLSKQGA-TECERWLEKPVIALGVFLMLVSLAG 1038
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
GAC R ++L+ LYL+VMF++I +L F IFA+VVT+KG+G V NRGY +Y L DYS
Sbjct: 1039 LVGACCRVSWLLWLYLLVMFVLIVLLFAFTIFAFVVTNKGAGEVVSNRGYKEYRLGDYSN 1218
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL++RV + W +I SC++ KVC + S + + FY L+++QSGCCKP
Sbjct: 1219 WLQKRVNNTKTWNRIRSCLQSGKVCTEF---QSKFLNDTVTEFYSENLSALQSGCCKPAE 1389
Query: 181 DCGYIYQNETVWNLGSGLMS-ANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
+C + Y N T W + + + +NPDC W+ND LC+ C SCKAG+L +LK W++V+V
Sbjct: 1390 ECQFSYVNPTTWTKPTNVTNQSNPDCDAWNNDPTVLCFNCQSCKAGLLQNLKTDWKRVAV 1569
Query: 240 INIVVMIILVIVYIIAYAAYRNNKR 264
+NIV ++ L+IVY I A+RNN+R
Sbjct: 1570 VNIVFLVFLIIVYSIGCCAFRNNRR 1644
>TC214305 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (89%)
Length = 1003
Score = 269 bits (687), Expect = 1e-72
Identities = 128/265 (48%), Positives = 184/265 (69%), Gaps = 1/265 (0%)
Frame = +1
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
+R SN++IG+LNFLTFLLSIPIL G+WLS + T+C ++L+ P+I +GV +M+VSLAG
Sbjct: 70 VRFSNNVIGLLNFLTFLLSIPILVAGVWLSKQGA-TECERWLEKPVIALGVFLMLVSLAG 246
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
GAC R ++L+ LYL+VMF++I +L F IFA+VVT+KG+G V NRGY +Y L DYS
Sbjct: 247 LVGACCRVSWLLWLYLLVMFVLIVLLFAFTIFAFVVTNKGAGEVVSNRGYKEYRLGDYSN 426
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL+++V + W +I+SC+ KVC + S + + FY L+++QSGCCKP
Sbjct: 427 WLQKKVNNTKTWNRISSCLHSGKVCTEF---QSKFLNDTVTQFYTEHLSALQSGCCKPAE 597
Query: 181 DCGYIYQNETVWNLGSGLMS-ANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSV 239
+C + Y+N T W + S NPDC W+N+Q LC+ C SCKAG L + K W++V+V
Sbjct: 598 ECLFTYENSTSWTKPGNVTSYNNPDCDAWNNNQTVLCFNCQSCKAGFLQNFKTEWKRVAV 777
Query: 240 INIVVMIILVIVYIIAYAAYRNNKR 264
+NIV +++L+IVY I A+RNN+R
Sbjct: 778 VNIVFLVLLIIVYSIGCCAFRNNRR 852
>TC203788 similar to GB|AAL91270.1|19699320|AY090367 AT3g12090/T21B14_110
{Arabidopsis thaliana;} , partial (84%)
Length = 1321
Score = 258 bits (660), Expect = 2e-69
Identities = 126/280 (45%), Positives = 174/280 (62%)
Frame = +1
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN +IG LN T L SIPI+G G+W++ ++T C FLQ PL++IG ++VVSLAGF
Sbjct: 169 RFSNTVIGFLNLFTLLASIPIIGAGLWMAR--SSTTCENFLQTPLLVIGFVVLVVSLAGF 342
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSGW 121
GAC+ + LYLVVM +IA L+G IF + VT KG G V R Y +Y+LQDYS W
Sbjct: 343 IGACFHVACALWLYLVVMLFLIAALMGLTIFGFGVTSKGGGVEVPGRVYKEYHLQDYSPW 522
Query: 122 LEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTD 181
L +R+ YW I C+ SK C K+ P D + R ++ +QSGCCKPPT
Sbjct: 523 LRKRIQDPRYWNTIRGCILGSKTCEKLASW------TPLD-YMQRDMSPIQSGCCKPPTA 681
Query: 182 CGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVIN 241
C Y N+ + +M+ +PDC +W+N LCY CDSCKAGVL ++ +W K+SV+
Sbjct: 682 CTY--------NVATTMMTQDPDCYRWNNAPNLLCYECDSCKAGVLEDIRGNWHKLSVLT 837
Query: 242 IVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKSHP 281
+ ++++L+ +Y I A+RN +R + D PYGE RMTK P
Sbjct: 838 VTMLVLLIGIYSIGCCAFRNTRRAETDYPYGENRMTKVRP 957
>TC228548 similar to GB|AAP13420.1|30023774|BT006312 At3g45600 {Arabidopsis
thaliana;} , partial (46%)
Length = 690
Score = 234 bits (598), Expect = 3e-62
Identities = 108/130 (83%), Positives = 117/130 (89%)
Frame = +1
Query: 157 PEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLC 216
PE D+FY+R LT +QSGCCKPPTDCGY+YQNETVW GSGLM ANPDC++WSNDQ LC
Sbjct: 1 PETPDMFYIRHLTPIQSGCCKPPTDCGYVYQNETVWIPGSGLMGANPDCTRWSNDQEQLC 180
Query: 217 YRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARM 276
Y CDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNN++MDNDEPYGEARM
Sbjct: 181 YACDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNRKMDNDEPYGEARM 360
Query: 277 TKSHPSHFQL 286
TK+ PS F L
Sbjct: 361 TKAQPSAFHL 390
>TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associated
protein-like protein, partial (30%)
Length = 1116
Score = 233 bits (593), Expect = 1e-61
Identities = 119/287 (41%), Positives = 177/287 (61%), Gaps = 1/287 (0%)
Frame = +1
Query: 1 MRTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAG 60
M SN++ VLN + L SIPI+ GIWL+S+ +N +C+ +WP++IIG+ I++VSLAG
Sbjct: 13 MGVSNNITAVLNLIATLASIPIIASGIWLASKPDN-ECVANFRWPIVIIGILILLVSLAG 189
Query: 61 FAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRGYMDYYLQDYSG 120
F GA + L+ LYL M L+IA+L+ ++FA+VVT V RGY +Y L +S
Sbjct: 190 FVGAYWNKQGLLALYLFCMALLIALLLLVLVFAFVVTRPDGAYDVPGRGYKEYRLHGFSS 369
Query: 121 WLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPT 180
WL V W KI C+ S VC+K+ + AD F+ ++ +QSGCCKPPT
Sbjct: 370 WLRNHVTGSGSWQKIRPCLAASDVCSKLTQDYIT-----ADQFFNSHISPLQSGCCKPPT 534
Query: 181 DCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVI 240
CGY Y N +W M A+ DC W+NDQ LCY C++CKAG+L +L+K WRK ++I
Sbjct: 535 ACGYNYVNPILWTNPVNPM-ADSDCYLWNNDQNQLCYNCNACKAGLLGNLRKEWRKANII 711
Query: 241 NIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARM-TKSHPSHFQL 286
IV +++L+ VY+IA +A+RN + + + Y + + ++ P H Q+
Sbjct: 712 LIVAVVVLIWVYVIACSAFRNAQTENLFDRYKQGWV*SRFPPQHLQV 852
>TC230196 weakly similar to UP|Q9SUD4 (Q9SUD4) Senescence-associated
protein-like, partial (37%)
Length = 662
Score = 132 bits (333), Expect = 1e-31
Identities = 54/126 (42%), Positives = 82/126 (64%)
Frame = +2
Query: 139 VRDSKVCAKMGRVDSNGIPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETVWNLGSGL 198
++ +K+C K +N + A FY L+++QSGCCKP DC + YQ +VWN G+
Sbjct: 8 LQSAKLCDKFETQFAN---DSAQQFYAENLSALQSGCCKPSNDCNFAYQGPSVWNKTDGV 178
Query: 199 MSANPDCSKWSNDQGFLCYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAA 258
+NPDC+ W ND LC+ C+SCKAG L +LK W+KV+++N++ ++ L+IVY + A
Sbjct: 179 NHSNPDCNAWDNDSNVLCFNCESCKAGFLQNLKTDWKKVTIVNVIFLVFLIIVYSVGCCA 358
Query: 259 YRNNKR 264
+RNN R
Sbjct: 359 FRNNMR 376
>BM178314 weakly similar to PIR|T02906|T029 senescence-associated protein
homolog T13J8.160 - Arabidopsis thaliana, partial (37%)
Length = 426
Score = 111 bits (278), Expect = 3e-25
Identities = 56/101 (55%), Positives = 78/101 (76%)
Frame = +1
Query: 2 RTSNHLIGVLNFLTFLLSIPILGGGIWLSSRANNTDCLKFLQWPLIIIGVSIMVVSLAGF 61
R SN L+G+LN L+FLLSIPIL G+WL +A T+C ++L+ PLI++GV ++VVSLAG
Sbjct: 124 RLSNSLVGLLNLLSFLLSIPILVTGVWLHKQAE-TECERWLEKPLIVLGVFLLVVSLAGL 300
Query: 62 AGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSG 102
GAC R + L+ LYL VMF++I V+ F +FA+VVT+KG+G
Sbjct: 301 LGACCRLSCLLWLYLFVMFILIIVVSAFTVFAFVVTNKGAG 423
>AW781408 similar to GP|19699320|gb| AT3g12090/T21B14_110 {Arabidopsis
thaliana}, partial (23%)
Length = 367
Score = 103 bits (256), Expect = 1e-22
Identities = 46/108 (42%), Positives = 67/108 (61%)
Frame = +2
Query: 41 FLQWPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKG 100
F Q PL++IG ++V+SLAGF GAC+ + + +YLV+M +IA L+ F +F +VVT++G
Sbjct: 5 FFQTPLLVIGFVVLVISLAGFIGACFHVAWALWVYLVIMVFIIAALLWFAVFGFVVTEQG 184
Query: 101 SGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKM 148
G V + + Y L YS WL R+ YW I SC+ S CAK+
Sbjct: 185 GGVEVPVKAFK*YCLDRYSPWLRNRIKDPQYWSSIMSCIWGSNTCAKL 328
>BI471574
Length = 422
Score = 100 bits (250), Expect = 6e-22
Identities = 56/166 (33%), Positives = 81/166 (48%)
Frame = +1
Query: 96 VTDKGSGRRVLNRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAKMGRVDSNG 155
VT V RGY +Y L +S WL V W KI C+ S VC+K+ +
Sbjct: 1 VTRPDGAYDVPGRGYKEYRLHGFSSWLRNHVTGSGSWQKIRPCLAASDVCSKLTQDYIT- 177
Query: 156 IPEPADVFYLRKLTSVQSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFL 215
AD F+ ++ +QS DC W+NDQ L
Sbjct: 178 ----ADQFFNSHISPLQS------------------------------DCYLWNNDQNQL 255
Query: 216 CYRCDSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRN 261
CY C++CKAG+L +L+K WRK ++I IV +++L+ VY+IA +A+RN
Sbjct: 256 CYNCNACKAGLLGNLRKEWRKANIILIVAVVVLIWVYVIACSAFRN 393
>CD392123
Length = 601
Score = 71.2 bits (173), Expect = 5e-13
Identities = 29/62 (46%), Positives = 44/62 (70%)
Frame = -2
Query: 220 DSCKAGVLASLKKSWRKVSVINIVVMIILVIVYIIAYAAYRNNKRMDNDEPYGEARMTKS 279
DSCKAGVL ++ +W K+SV+ + ++++L+ +Y I A+RN +R + D PYGE RMTK
Sbjct: 381 DSCKAGVLEDIRGNWHKLSVLTVTMLVLLIGIYSIGCCAFRNTRRAETDYPYGENRMTKV 202
Query: 280 HP 281
P
Sbjct: 201 RP 196
>BE210525
Length = 381
Score = 70.1 bits (170), Expect = 1e-12
Identities = 28/65 (43%), Positives = 42/65 (64%)
Frame = +1
Query: 167 KLTSVQSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSCKAGV 226
KL+ +++GCC+PP+ CGY N + ++L +S N DC ++ N + CY CDSCKAGV
Sbjct: 187 KLSPIEAGCCRPPSQCGYPAVNASYYDLTFHPVSPNNDCKRYKNSRAIKCYDCDSCKAGV 366
Query: 227 LASLK 231
+K
Sbjct: 367 AQYMK 381
>AW780642
Length = 142
Score = 54.7 bits (130), Expect = 5e-08
Identities = 23/48 (47%), Positives = 27/48 (55%)
Frame = +2
Query: 175 CCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYRCDSC 222
CCKPPT CGY Y N +W + A+ C WSND LCY D+C
Sbjct: 2 CCKPPTACGYYYVNPILWT-NPVIPMADSYCYLWSNDLNHLCYHSDAC 142
>AW780677
Length = 166
Score = 33.5 bits (75), Expect = 0.11
Identities = 21/54 (38%), Positives = 25/54 (45%), Gaps = 2/54 (3%)
Frame = +2
Query: 171 VQSGCCKPPTDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYR--CDSC 222
+QS C KP T CGYI N +W M A+ D W+ D L Y C C
Sbjct: 8 LQSRCSKPTTACGYI*FNPILWTNPVNPM-ADSDVYLWNIDHNQLFYT**CVQC 166
>TC227413 similar to UP|Q7PS25 (Q7PS25) ENSANGP00000020753 (Fragment),
partial (38%)
Length = 1408
Score = 29.3 bits (64), Expect = 2.1
Identities = 20/62 (32%), Positives = 26/62 (41%), Gaps = 2/62 (3%)
Frame = +3
Query: 175 CCKPP-TDCGYIYQNETVWNLGSGLMSANPDCSKWSNDQGFLCYR-CDSCKAGVLASLKK 232
CC P CG+ + E SG P CSKW + LC + C C + L + K
Sbjct: 135 CCVPH*CYCGFCHCEE----FDSGSEGQKPSCSKWIYKRN*LCKKFCQGC-S*QLPAFPK 299
Query: 233 SW 234
W
Sbjct: 300 EW 305
>TC211402
Length = 508
Score = 28.5 bits (62), Expect = 3.7
Identities = 10/31 (32%), Positives = 22/31 (70%)
Frame = +1
Query: 27 IWLSSRANNTDCLKFLQWPLIIIGVSIMVVS 57
+WLSS +++T KF PL+++ + ++++S
Sbjct: 46 LWLSSGSDHTSMAKFGLSPLLVLSILLLIMS 138
>AI460424
Length = 285
Score = 28.1 bits (61), Expect = 4.8
Identities = 15/45 (33%), Positives = 22/45 (48%)
Frame = -3
Query: 65 CYRNTFLMRLYLVVMFLVIAVLIGFIIFAYVVTDKGSGRRVLNRG 109
C RN RL V FL + + +FAY+ KG+G+ + G
Sbjct: 172 CQRNCEWTRLSEEVKFLPVHYINTRFLFAYIFLTKGNGQLKITEG 38
>TC213340
Length = 501
Score = 28.1 bits (61), Expect = 4.8
Identities = 14/59 (23%), Positives = 30/59 (50%), Gaps = 5/59 (8%)
Frame = +1
Query: 44 WPLIIIGVSIMVVSLAGFAGACYRNTFLMRLYLVVMFLV-----IAVLIGFIIFAYVVT 97
WP + + + +S F +C+ N ++L+ ++ FLV +++LI F V++
Sbjct: 118 WPFLHMSKTNYSISYRPFLNSCHTNISFIQLFCILAFLVWPFIYLSILISITHFFLVLS 294
>TC216816 weakly similar to UP|KHLX_HUMAN (Q9Y2M5) Kelch-like protein X,
partial (5%)
Length = 1610
Score = 28.1 bits (61), Expect = 4.8
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = -2
Query: 107 NRGYMDYYLQDYSGWLEERVASHSYWGKIASCVRDSKVCAK 147
NR + +YLQ + ++ VA +W + AS + S C +
Sbjct: 304 NRTHYPHYLQGHKEQMQHYVAKSHFWSQEASVMASSHHCVQ 182
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.140 0.449
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,112,254
Number of Sequences: 63676
Number of extensions: 264221
Number of successful extensions: 1600
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1576
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1584
length of query: 286
length of database: 12,639,632
effective HSP length: 96
effective length of query: 190
effective length of database: 6,526,736
effective search space: 1240079840
effective search space used: 1240079840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0246c.3


