

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.8
(334 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
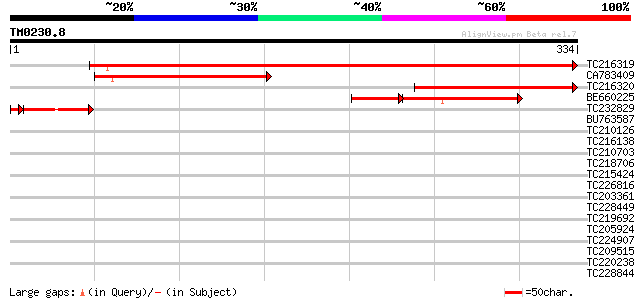
Score E
Sequences producing significant alignments: (bits) Value
TC216319 516 e-147
CA783409 184 5e-47
TC216320 177 5e-45
BE660225 similar to GP|13872875|dbj P0434B04.3 {Oryza sativa (ja... 103 3e-35
TC232829 similar to UP|SYN2_HUMAN (Q92777) Synapsin II, partial ... 71 3e-13
BU763587 32 0.31
TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C2... 32 0.53
TC216138 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37... 30 1.5
TC210703 similar to UP|Q96397 (Q96397) LRG5 protein, partial (4%) 30 1.5
TC218706 similar to UP|Q9FFC0 (Q9FFC0) Histone H2B like protein,... 29 2.6
TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%) 29 2.6
TC226816 similar to GB|AAA34858.1|172136|YSCPFK 6-phosphofructo-... 29 3.4
TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 28 4.4
TC228449 similar to GB|AAR99134.1|41058179|BT011476 RE11923p {Dr... 28 4.4
TC219692 similar to UP|Q6TVN2 (Q6TVN2) ORF088 virion core protei... 28 5.8
TC205924 homologue to UP|Q70I19 (Q70I19) TCP-domain protein, par... 28 5.8
TC224907 weakly similar to UP|Q6Z8A6 (Q6Z8A6) Zinc finger (C3HC4... 28 7.6
TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6,... 28 7.6
TC220238 UP|Q84VH5 (Q84VH5) Envelope-like protein (Fragment), co... 28 7.6
TC228844 weakly similar to GB|AAO44030.1|28466843|BT004764 At3g5... 27 9.9
>TC216319
Length = 1306
Score = 516 bits (1328), Expect = e-147
Identities = 260/289 (89%), Positives = 276/289 (94%), Gaps = 2/289 (0%)
Frame = +2
Query: 48 EINDLFKMG--KKKNERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLL 105
EI DLFK+G KKKNERSPAEIA LVE V+AELEVTAEEDAELNRQGKPA+NKLKKL LL
Sbjct: 2 EIKDLFKIGQKKKKNERSPAEIAYLVETVMAELEVTAEEDAELNRQGKPAVNKLKKLNLL 181
Query: 106 TEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQIDRR 165
TEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRT +LKILNDFPIDLEQ DRR
Sbjct: 182 TEVLSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTEVLKILNDFPIDLEQYDRR 361
Query: 166 EQLKRSGLGKVIMFLSKSDEEINVNRKLTKELVDKWSRPIFNKSTRFEDMRNIEDERAPF 225
EQLK+SGLGKVIMFLSKSDEEINVNRKL KELVDKWSRPIFNKSTRFEDMRN+ED+R P+
Sbjct: 362 EQLKKSGLGKVIMFLSKSDEEINVNRKLAKELVDKWSRPIFNKSTRFEDMRNVEDDRVPY 541
Query: 226 RRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDP 285
RRPSVKKPANKA GM+SRDSDLDL+L QPRSGQSSSRQHASRPEATP+DFV+RPQSK+DP
Sbjct: 542 RRPSVKKPANKAAGMESRDSDLDLELSQPRSGQSSSRQHASRPEATPLDFVVRPQSKIDP 721
Query: 286 EEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
EE+RARAKQA+ DQQRMKMNKKLQQLRAPKK+QLQATKLSVEGRGMIKY
Sbjct: 722 EEIRARAKQAAQDQQRMKMNKKLQQLRAPKKKQLQATKLSVEGRGMIKY 868
>CA783409
Length = 430
Score = 184 bits (467), Expect = 5e-47
Identities = 96/106 (90%), Positives = 100/106 (93%), Gaps = 2/106 (1%)
Frame = +2
Query: 51 DLFKMGKKK--NERSPAEIALLVENVVAELEVTAEEDAELNRQGKPAINKLKKLPLLTEV 108
DLFK+GKKK NERSPAEIA LVE V+AELEVTAEEDAELNRQGKPAINKLKKL LLTEV
Sbjct: 5 DLFKIGKKKKKNERSPAEIAYLVETVMAELEVTAEEDAELNRQGKPAINKLKKLNLLTEV 184
Query: 109 LSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRTAILKILND 154
LSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIR+ +LKILND
Sbjct: 185 LSKKQLQLEFLDHGVLTLLKNWLEPLPDGSLPNINIRSEVLKILND 322
>TC216320
Length = 597
Score = 177 bits (450), Expect = 5e-45
Identities = 86/96 (89%), Positives = 94/96 (97%)
Frame = +3
Query: 239 GMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHD 298
GM+SRDSDLDL+L QPRSGQSSSRQHASRPEATP+DFV+RPQSK+DPEE+RARAKQA+ D
Sbjct: 9 GMESRDSDLDLELSQPRSGQSSSRQHASRPEATPLDFVVRPQSKIDPEEIRARAKQAAQD 188
Query: 299 QQRMKMNKKLQQLRAPKKRQLQATKLSVEGRGMIKY 334
QQRMKMNKKLQQLRAPKK+QLQATKLSVEGRGMIKY
Sbjct: 189 QQRMKMNKKLQQLRAPKKKQLQATKLSVEGRGMIKY 296
>BE660225 similar to GP|13872875|dbj P0434B04.3 {Oryza sativa (japonica
cultivar-group)}, partial (15%)
Length = 612
Score = 103 bits (258), Expect(2) = 3e-35
Identities = 57/103 (55%), Positives = 64/103 (61%), Gaps = 32/103 (31%)
Frame = +3
Query: 232 KPANKAPGMQSRDSDLDLDLPQ--------------------------------PRSGQS 259
+PANKA GM+SRDSDLDL+L Q PRSGQS
Sbjct: 225 RPANKAAGMESRDSDLDLELSQ*GLNGIYELVVEHCGLSNH*NILYVKFFH*CRPRSGQS 404
Query: 260 SSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRM 302
SSRQHASRPEATP+DFV+RPQSK+DPEE+RARAKQA RM
Sbjct: 405 SSRQHASRPEATPLDFVVRPQSKIDPEEIRARAKQADKISSRM 533
Score = 62.4 bits (150), Expect(2) = 3e-35
Identities = 27/31 (87%), Positives = 30/31 (96%)
Frame = +1
Query: 202 SRPIFNKSTRFEDMRNIEDERAPFRRPSVKK 232
SRPIFNKSTRFEDMRN+ED+R P+RRPSVKK
Sbjct: 43 SRPIFNKSTRFEDMRNVEDDRVPYRRPSVKK 135
>TC232829 similar to UP|SYN2_HUMAN (Q92777) Synapsin II, partial (3%)
Length = 425
Score = 71.2 bits (173), Expect(2) = 3e-13
Identities = 32/41 (78%), Positives = 38/41 (92%)
Frame = -3
Query: 9 LDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEI 49
+DDDNFIDDTGVEP +YG+ +EP SPG+APQAEEGEED+EI
Sbjct: 120 VDDDNFIDDTGVEPAYYGS-DEPRSPGDAPQAEEGEEDEEI 1
Score = 21.2 bits (43), Expect(2) = 3e-13
Identities = 8/8 (100%), Positives = 8/8 (100%)
Frame = -2
Query: 1 DDQEGVRN 8
DDQEGVRN
Sbjct: 145 DDQEGVRN 122
>BU763587
Length = 368
Score = 32.3 bits (72), Expect = 0.31
Identities = 12/39 (30%), Positives = 21/39 (53%)
Frame = -1
Query: 124 LTLLKNWLEPLPDGSLPNINIRTAILKILNDFPIDLEQI 162
L + W PLP S P++++ + L + FP+DL +
Sbjct: 179 LEWVSRWFLPLPPESPPSVDVNLSYLAFIGPFPVDLNPV 63
>TC210126 similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23), partial
(7%)
Length = 735
Score = 31.6 bits (70), Expect = 0.53
Identities = 17/52 (32%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Frame = +3
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNY-NEPSSPGEAPQAEEGEEDDEIND 51
DD++G + +D + DD G + F G +E P + P+A G DD+ +D
Sbjct: 345 DDKDGDEDDEDGHDQDDDGEDEEFSGEEGDEEGDPEDDPEANGGGSDDDDDD 500
>TC216138 similar to UP|LB37_ARATH (Q9FN11) LOB domain protein 37, partial
(50%)
Length = 1169
Score = 30.0 bits (66), Expect = 1.5
Identities = 16/52 (30%), Positives = 25/52 (47%)
Frame = -1
Query: 232 KPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKV 283
+P N+ G SR L + + P S ASRP+++ M + P+S V
Sbjct: 572 EPGNRKFGFGSRTLHLSVQVTSPSEASSVVGVGASRPKSSGMGLTVPPRSTV 417
>TC210703 similar to UP|Q96397 (Q96397) LRG5 protein, partial (4%)
Length = 491
Score = 30.0 bits (66), Expect = 1.5
Identities = 19/56 (33%), Positives = 28/56 (49%)
Frame = -2
Query: 29 NEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIALLVENVVAELEVTAEE 84
N ++P PQAEE +E D + G ++ ++ P L E VVA +EV E
Sbjct: 208 NHIATPSNLPQAEEEAWVEETLDSSEFGDRRLKKLPPPQLELEEVVVAAMEVAMLE 41
>TC218706 similar to UP|Q9FFC0 (Q9FFC0) Histone H2B like protein, partial
(94%)
Length = 583
Score = 29.3 bits (64), Expect = 2.6
Identities = 15/62 (24%), Positives = 33/62 (53%), Gaps = 1/62 (1%)
Frame = +1
Query: 226 RRPSVKKPANKAPGMQSRDSDLDLDLP-QPRSGQSSSRQHASRPEATPMDFVIRPQSKVD 284
++P+ KKPA KAP +++ + +P +P SG+ ++ + T ++ + +V
Sbjct: 73 KKPAEKKPAEKAPAEKTK---AEKKIPKEPASGEKKKKKRNKKSVETYKIYIFKVLKQVH 243
Query: 285 PE 286
P+
Sbjct: 244 PD 249
>TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%)
Length = 1030
Score = 29.3 bits (64), Expect = 2.6
Identities = 17/69 (24%), Positives = 31/69 (44%)
Frame = +3
Query: 262 RQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRAPKKRQLQA 321
+QH +P P + + +E + K+ ++QR + K QQL+ + LQ
Sbjct: 405 KQHKQQPPVPVKKMNNGPPGRAETDEEKRLRKKREFEKQRQE-EKHRQQLKESQNTVLQK 581
Query: 322 TKLSVEGRG 330
T + G+G
Sbjct: 582 THMLSSGKG 608
>TC226816 similar to GB|AAA34858.1|172136|YSCPFK 6-phosphofructo-2-kinase
{Saccharomyces cerevisiae;} , partial (3%)
Length = 679
Score = 28.9 bits (63), Expect = 3.4
Identities = 12/40 (30%), Positives = 20/40 (50%)
Frame = +3
Query: 27 NYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAE 66
N PS PG P+ + ++ DLF+ ++N+ AE
Sbjct: 402 NKTTPSEPGSVPEPSPTTQKKKLLDLFRESVRENQSDDAE 521
>TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 1090
Score = 28.5 bits (62), Expect = 4.4
Identities = 19/59 (32%), Positives = 25/59 (42%)
Frame = -1
Query: 1 DDQEGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKK 59
D+ E + DDD DD + G E E + EE EED+E L K+K
Sbjct: 412 DEDEDGDDQDDDEDDDDDDDDEEDEGGEEEDEDGAEDEENEEDEEDEEEEALQPPKKRK 236
>TC228449 similar to GB|AAR99134.1|41058179|BT011476 RE11923p {Drosophila
melanogaster;} , partial (3%)
Length = 1112
Score = 28.5 bits (62), Expect = 4.4
Identities = 27/100 (27%), Positives = 42/100 (42%), Gaps = 6/100 (6%)
Frame = +1
Query: 224 PFRRPSVKKPANKAPGMQSRDSDLDLDLPQPRSGQSSSRQHASR--PEATPMDFVIRPQS 281
P RR + KP+ P S S P PRS SSSR +S + +P P+S
Sbjct: 259 PRRRRPLSKPS---PSPLSPTSSRPTTTPSPRSSPSSSRSSSSEKTTKRSPSSSASSPRS 429
Query: 282 KVDPEEV---RARAKQASHDQQRMKMNKKLQ-QLRAPKKR 317
P RA ++ + + R ++ + R+P +R
Sbjct: 430 PTPPTGSSWRRASPRRPATSRPRAATTRRSSPRTRSPSRR 549
>TC219692 similar to UP|Q6TVN2 (Q6TVN2) ORF088 virion core protein, partial
(9%)
Length = 1220
Score = 28.1 bits (61), Expect = 5.8
Identities = 23/91 (25%), Positives = 42/91 (45%), Gaps = 5/91 (5%)
Frame = -3
Query: 4 EGVRNLDDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEIN-----DLFKMGKK 58
E ++++ +D V+ G E + + P E ++DDE+N DL
Sbjct: 306 EAAAGVNEN*GVDALEVDKNIEGELAEVTETVDVPPKENPDDDDEVNVEPNGDLLGCQLD 127
Query: 59 KNERSPAEIALLVENVVAELEVTAEEDAELN 89
N+ E+ L+E E+TAEE+A+++
Sbjct: 126 PNK---VELGPLLELT----ELTAEEEADVS 55
>TC205924 homologue to UP|Q70I19 (Q70I19) TCP-domain protein, partial (17%)
Length = 704
Score = 28.1 bits (61), Expect = 5.8
Identities = 25/82 (30%), Positives = 32/82 (38%)
Frame = +2
Query: 249 LDLPQPRSGQSSSRQHASRPEATPMDFVIRPQSKVDPEEVRARAKQASHDQQRMKMNKKL 308
L LPQ + G +++ A DF I V A K+ S KK
Sbjct: 209 LSLPQQQQGNTNNNNMGENKPAEVKDFQI----------VVAENKEES---------KKQ 331
Query: 309 QQLRAPKKRQLQATKLSVEGRG 330
QQ APK+ + VEGRG
Sbjct: 332 QQQLAPKRSSNKDRHTKVEGRG 397
>TC224907 weakly similar to UP|Q6Z8A6 (Q6Z8A6) Zinc finger (C3HC4-type RING
finger)-like, partial (12%)
Length = 1685
Score = 27.7 bits (60), Expect = 7.6
Identities = 19/51 (37%), Positives = 25/51 (48%), Gaps = 5/51 (9%)
Frame = -2
Query: 223 APFRRPS--VKKPANKAPGMQSRDSDLDLDLPQPR---SGQSSSRQHASRP 268
+P RPS ++ P+++AP SR DLP P SSSR S P
Sbjct: 1609 SPSHRPSTELQTPSSRAPSYPSRRRRRSPDLPAPSISPLSNSSSRSSPSSP 1457
>TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6, Srg6
(Stress responsive gene 6 protein, Srg6), partial (55%)
Length = 1386
Score = 27.7 bits (60), Expect = 7.6
Identities = 19/87 (21%), Positives = 43/87 (48%), Gaps = 3/87 (3%)
Frame = +2
Query: 10 DDDNFIDDTGVEPGFYGNYNEPSSPGEAPQAEEGEEDDEINDLFKMGKKKNERSPAEIAL 69
+DDN+ ++ + F ++++ + P EE E+ ++ +D +M KKK P +
Sbjct: 107 EDDNYQEEPEIADEFDSDFDQ-----DEPVPEEEEDPNKNDDDERMHKKKRLIFPGKTLA 271
Query: 70 ---LVENVVAELEVTAEEDAELNRQGK 93
+ +++LE + +E+ + GK
Sbjct: 272 KKKKKKKTISKLESSPKENEDEEHSGK 352
>TC220238 UP|Q84VH5 (Q84VH5) Envelope-like protein (Fragment), complete
Length = 2128
Score = 27.7 bits (60), Expect = 7.6
Identities = 25/103 (24%), Positives = 46/103 (44%), Gaps = 10/103 (9%)
Frame = +1
Query: 182 KSDEEINVNRK---LTKELVDKWSRPIFNKSTRFEDMRNIEDERAPF-----RRPSVKKP 233
+SDEE N+ + + L + + +S R + M + + P RR V P
Sbjct: 712 ESDEEPIANKLAPGIAERLQSRKGKTPIKRSGRIKTMA--QKKSTPITPTTSRRSKVAIP 885
Query: 234 ANKAPGMQSRDSD--LDLDLPQPRSGQSSSRQHASRPEATPMD 274
+ K + S DSD ++LD+P + ++S ++ P+D
Sbjct: 886 SKKRKEISSSDSDDDVELDVPDIKRAKTSGKKVPGNVPDAPLD 1014
>TC228844 weakly similar to GB|AAO44030.1|28466843|BT004764 At3g56220
{Arabidopsis thaliana;} , partial (40%)
Length = 718
Score = 27.3 bits (59), Expect = 9.9
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = +2
Query: 278 RPQSKVDPEEVRARAKQASHDQQRMKMNKKLQQLRA 313
+P++K++ EE R+ + + M +KLQQLR+
Sbjct: 20 KPKTKINLEEKVGRSMDSRRRGKNASMQRKLQQLRS 127
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.133 0.367
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,810,415
Number of Sequences: 63676
Number of extensions: 126423
Number of successful extensions: 658
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 644
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 656
length of query: 334
length of database: 12,639,632
effective HSP length: 98
effective length of query: 236
effective length of database: 6,399,384
effective search space: 1510254624
effective search space used: 1510254624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0230.8


