

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0230.4
(606 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
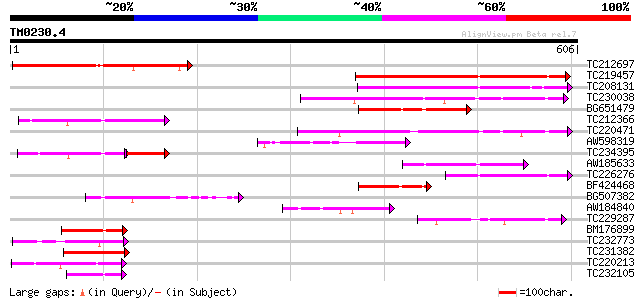
Score E
Sequences producing significant alignments: (bits) Value
TC212697 weakly similar to UP|O13633 (O13633) HLJ1 PROTEIN (SPBC... 233 2e-61
TC219457 similar to GB|AAN46891.1|24111445|BT001092 At2g05250/F5... 167 1e-41
TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein... 163 2e-40
TC230038 115 4e-26
BG651479 97 3e-20
TC212366 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein (At2g01... 97 3e-20
TC220471 93 3e-19
AW598319 86 5e-17
TC234395 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein (At2g01... 76 5e-14
AW185633 71 1e-12
TC226276 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein... 69 6e-12
BF424468 66 5e-11
BG507382 66 5e-11
AW184840 62 7e-10
TC229287 similar to UP|Q8RYF9 (Q8RYF9) Heat shock protein-like, ... 60 3e-09
BM176899 similar to GP|27817868|dbj OJ1081_B12.17 {Oryza sativa ... 55 1e-07
TC232773 similar to UP|Q9S729 (Q9S729) GlsA, partial (4%) 55 1e-07
TC231382 weakly similar to GB|AAP21357.1|30102878|BT006549 At1g5... 54 2e-07
TC220213 similar to UP|Q7XYV0 (Q7XYV0) P58IPK, partial (38%) 54 2e-07
TC232105 similar to UP|Q9FEW7 (Q9FEW7) DnaJ like protein, partia... 53 4e-07
>TC212697 weakly similar to UP|O13633 (O13633) HLJ1 PROTEIN (SPBC17A3.05c
protein) (Pi041 protein), partial (10%)
Length = 741
Score = 233 bits (594), Expect = 2e-61
Identities = 121/201 (60%), Positives = 147/201 (72%), Gaps = 9/201 (4%)
Frame = -3
Query: 4 SKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATDWY 63
++ E EALRLK +AESK + SNN KSALKYA RA RL P L G+ E V +L++L+A DWY
Sbjct: 598 AEAESEALRLKAMAESKFKASNNAKSALKYANRAHRLCPHLAGVPETVAALSVLAAPDWY 419
Query: 64 TVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKG 123
LG EPFA+S+ +R+QYKKL+LLLHPDK N HV ASEEAFKL+GEAFR LSD ++
Sbjct: 418 RALGAEPFASSSVIRRQYKKLALLLHPDK-NPHV--ASEEAFKLLGEAFRFLSDRNRRRE 248
Query: 124 YDAELRKK------EAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSDG 177
YDAELR+K E+ TFWTACS CRLLHQFERRY+G LVCP+C K F AVEAV SD
Sbjct: 247 YDAELRRKIEAAESESETFWTACSTCRLLHQFERRYLGQELVCPSCEKGFRAVEAVQSDD 68
Query: 178 SSD---EGEKVGVRRNEKREV 195
D ++ ++ EKRE+
Sbjct: 67 DGDVRVRSRRLKLKEMEKREI 5
>TC219457 similar to GB|AAN46891.1|24111445|BT001092 At2g05250/F5G3.15
{Arabidopsis thaliana;} , partial (31%)
Length = 1054
Score = 167 bits (424), Expect = 1e-41
Identities = 91/231 (39%), Positives = 140/231 (60%), Gaps = 1/231 (0%)
Frame = +3
Query: 370 MAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDL 429
+ V DSDF+DFDKDR E F+ Q+WA YD++DGMPR Y +I E VS NPF++ IS+L
Sbjct: 48 ITVPDSDFHDFDKDRSEECFRPKQIWALYDEEDGMPRLYCMIREVVSVNPFKIHISYLSS 227
Query: 430 QSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAR-EVYKIYPRKGS 488
+++++ V+ GF CG F+ D+++ VNIFSHV+ ++ R +IYPR G
Sbjct: 228 KTDSEFGSVNWLDSGFTKSCGNFRAFNSDAVDQVNIFSHVLSKEKAGRGGCVRIYPRSGD 407
Query: 489 VWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDT 548
+WA+Y + D + R DE + ++V L +Y+E G+ ++ L K+ G+KTV+ + +T
Sbjct: 408 IWAVYRNWSPDWN-RSTPDEVRHQYEMVEVLDDYSEELGVCVSPLIKLAGFKTVY-QSNT 581
Query: 549 GSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLL 599
I+++ + M SHQ+P+ E L E CW+LDPA+ P LL
Sbjct: 582 DKSTIKWIPRREMLRFSHQVPSWLLK--GEASNLPERCWDLDPAATPDELL 728
>TC208131 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like,
partial (32%)
Length = 1371
Score = 163 bits (413), Expect = 2e-40
Identities = 96/230 (41%), Positives = 133/230 (57%)
Frame = +3
Query: 372 VVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQS 431
V D DF++FD DR E SF + QVWAAYDDDDGMPR+YA I + +S PF++RISWL+ +S
Sbjct: 129 VPDPDFHNFDLDRDENSFAEDQVWAAYDDDDGMPRYYAKIHKVISMKPFKMRISWLNSRS 308
Query: 432 NADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWA 491
N++ + GF+ CG F+ + + S+N FSH V + R V +I+P KG VWA
Sbjct: 309 NSELGPIDWVGSGFYKTCGDFRTGKHEITESLNSFSHKVRWTKGTRGVVRIFPGKGEVWA 488
Query: 492 LYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTVFKRQDTGSH 551
LY + D + H DE D+V L +++E G+ + L KV G++TVF+R
Sbjct: 489 LYRNWSPDWN-EHTPDEVIHKYDMVEVLEDFDEEQGILVTPLVKVAGFRTVFQRHMDCDQ 665
Query: 552 AIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLLTI 601
R L K+ M+ SHQ+P E + C ELDPA+ P LL I
Sbjct: 666 ERRIL-KEEMFQFSHQVP-NYLLTGQEADNAPKGCRELDPAATPLDLLQI 809
>TC230038
Length = 1229
Score = 115 bits (289), Expect = 4e-26
Identities = 92/308 (29%), Positives = 143/308 (45%), Gaps = 21/308 (6%)
Frame = +3
Query: 311 KLPIKENA-VKSKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEECGDANGGEL-- 367
++ + EN+ VK R S GE + + + +AS + LE N +
Sbjct: 63 EIAVPENSDVKVGRSSSGGENTRPSNRSEPLMTSEGDASIPKVNLERSNLATENKDSVDD 242
Query: 368 ------------EVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETV 415
E + V D+ F+DFD R F+ GQ+WA Y D+DG+P++Y I +
Sbjct: 243 SDNCCAPPESSPEAINVPDTQFFDFDGGRALEKFQIGQIWAFYSDEDGLPKYYGQIKKIE 422
Query: 416 SANPFEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSV----NIFSHVVD 471
++ E+ + WL + I +K I CGRFKV SV + SH V
Sbjct: 423 TSPDLELHVYWLTCCWLPENTIKWEDK-DILISCGRFKVNETHDFLSVYSTTSCVSHQVH 599
Query: 472 CDRVAR-EVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSM 530
D V + + Y I+PRKG VWALY + T + E + C +V + ++ +++
Sbjct: 600 ADAVGKNKNYAIFPRKGDVWALYRKWT----NKMKCFEMENCEYDIVEVVEETDL-FINV 764
Query: 531 AYLEKVDGYKTVFK-RQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWEL 589
LE V GY +VF+ + + GS + + + SHQIPA K + E L+ WEL
Sbjct: 765 LVLEFVSGYTSVFRGKSNEGSSVNLRIPRKELLRFSHQIPAFK---LTEEHGNLKGFWEL 935
Query: 590 DPASLPSY 597
DP +LP +
Sbjct: 936 DPGALPMH 959
>BG651479
Length = 392
Score = 96.7 bits (239), Expect = 3e-20
Identities = 47/122 (38%), Positives = 74/122 (60%), Gaps = 2/122 (1%)
Frame = +1
Query: 374 DSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNA 433
D++F DFDK + + F GQ+WA YD +GMPR YALI + +S F ++I W + +
Sbjct: 19 DAEFSDFDKGKNKECFTAGQIWAIYDTSEGMPRFYALIRKVLSPG-FRLQIIWFEPHPDC 195
Query: 434 DGKI--VSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREVYKIYPRKGSVWA 491
+I V+ E + CG++K++ D +FSH V C++++R +K+YPRKG WA
Sbjct: 196 KDEINWVNEE---MPVACGKYKLSDIDITEDHLMFSHPVLCEKISRNTFKVYPRKGETWA 366
Query: 492 LY 493
L+
Sbjct: 367 LF 372
>TC212366 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein
(At2g01710/T8O11.12), partial (50%)
Length = 701
Score = 96.7 bits (239), Expect = 3e-20
Identities = 63/172 (36%), Positives = 91/172 (52%), Gaps = 11/172 (6%)
Frame = +1
Query: 10 ALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSAT--------- 60
A RL + E LQ S + S+ +A AQ P LEG +++ + +L A
Sbjct: 1 AERLLAIGEKLLQ-SRDLSSSRDFAILAQEAEPLLEGSDQILAIVEVLLAAEKPITNDHL 177
Query: 61 DWYTVLGVEPFANS-NAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
DWY +L V+ + ++KQY++L LLLHPDKN +A + AFKLV +A+ VLSD
Sbjct: 178 DWYAILQVDRTCQDLDLIKKQYRRLGLLLHPDKNPFSLA---DHAFKLVSDAWAVLSDPV 348
Query: 120 LKKGYDAELRKKEAP-TFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV 170
K YD ++ P +FWTAC C L+++ G L C NC +SF +
Sbjct: 349 QKAIYDRDVAGSVQPESFWTACPYCYFLYEYPAVCEGCCLRCQNCERSFHGL 504
>TC220471
Length = 962
Score = 93.2 bits (230), Expect = 3e-19
Identities = 83/302 (27%), Positives = 131/302 (42%), Gaps = 8/302 (2%)
Frame = +3
Query: 308 RTLKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGKR----KRLELEECGDAN 363
+T+K +K K E GE +K+ + K+ + K+ + N
Sbjct: 60 QTIKRKVKTPQKHEKNNYE-GEALKARKSPRDLSKKNAQGDAGEWTAGKKTDNHSSNSKN 236
Query: 364 GGELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVR 423
+ V + Y F K++ E F+ GQ+WA Y D D MP YA I F ++
Sbjct: 237 VKVSNIPQSVGASCYGFKKEKSEEMFQCGQIWAIYGDRDHMPDTYAQIRMIECTPNFRLQ 416
Query: 424 ISWLDLQSNADGKIVSREKMGFHIPCGRFKVAR-KDSINSVNIFSHVVDCDRVAREVYKI 482
+ L+ + I CG F V K + S++ FSH + + VA Y+I
Sbjct: 417 VYMLE-------PCPPPNDLKRTISCGTFSVKEAKLRMLSLSAFSHQLKAELVANNRYEI 575
Query: 483 YPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNGLSMAYLEKVDGYKTV 542
YPRKG +WALY + + +++G+ IV L + N+ + + L +T+
Sbjct: 576 YPRKGEIWALYKDQNYEQTS---SNQGRGECHIVEVLADNNK--SIQVVVLVPHGNSQTI 740
Query: 543 FKR---QDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLPSYLL 599
FK Q + + I L K+ + SHQIPA + L CWELDP+S+P +
Sbjct: 741 FKAPRIQRSKTGVIEILRKE-VGRFSHQIPAFQ----HSDNVHLRGCWELDPSSVPGSFI 905
Query: 600 TI 601
I
Sbjct: 906 PI 911
>AW598319
Length = 440
Score = 85.9 bits (211), Expect = 5e-17
Identities = 63/168 (37%), Positives = 84/168 (49%), Gaps = 4/168 (2%)
Frame = +3
Query: 265 QSQVKR----KVQQEKVKDKEEEWGRTDKRSSRKRVENKRGLEIGEVRTLKLPIKENAVK 320
Q Q+K K ++E VK K +EW K SS +K E+R + N VK
Sbjct: 9 QHQIKNILVEKARKEIVK-KLDEW----KASSASNNLDKSKNTDTEIREKGKEREGNVVK 173
Query: 321 SKRGSEVGEERSLKKNVKLAIKEKPEASGKRKRLELEECGDANGGELEVMAVVDSDFYDF 380
G++V + ++ K A E PE G M V D DF+DF
Sbjct: 174 P--GAQVVDSETVNKKCFSADLE-PELPGSLS-----------------MNVPDPDFHDF 293
Query: 381 DKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLD 428
D DR E +F + QVWAAYD+DDGMPR+Y LI + +S NP +RISWL+
Sbjct: 294 DGDRTENAFGENQVWAAYDNDDGMPRYYCLIHDVISKNPLNMRISWLN 437
>TC234395 similar to UP|Q9ZU99 (Q9ZU99) Expressed protein
(At2g01710/T8O11.12), partial (48%)
Length = 760
Score = 75.9 bits (185), Expect = 5e-14
Identities = 51/129 (39%), Positives = 75/129 (57%), Gaps = 9/129 (6%)
Frame = +3
Query: 9 EALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATD------- 61
EA RL G+AE LQ + S ++A AQ P LEG +++ + +L A D
Sbjct: 54 EAERLLGIAEKLLQNRDLVGSR-EFAFLAQETEPLLEGSDQILAIVDVLLAADKRVNNHP 230
Query: 62 -WYTVLGVEPFANS-NAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSA 119
WY VL V+ ++ + ++KQY++L+LLLHPDK+ H A + AF+LV +A+ +LSD
Sbjct: 231 DWYAVLQVDRRSDDLDLIKKQYRRLALLLHPDKSRFHFA---DHAFQLVADAWALLSDPI 401
Query: 120 LKKGYDAEL 128
K YD EL
Sbjct: 402 KKSVYDKEL 428
Score = 48.5 bits (114), Expect = 8e-06
Identities = 20/46 (43%), Positives = 28/46 (60%)
Frame = +3
Query: 125 DAELRKKEAPTFWTACSACRLLHQFERRYVGHSLVCPNCNKSFEAV 170
D R++ + TFWTAC C L+++ R Y G L C NC++SF V
Sbjct: 582 DENSRRRRSSTFWTACPYCYRLYEYPRVYEGCCLRCQNCDRSFHGV 719
>AW185633
Length = 443
Score = 71.2 bits (173), Expect = 1e-12
Identities = 41/137 (29%), Positives = 72/137 (51%), Gaps = 2/137 (1%)
Frame = +1
Query: 420 FEVRISWLDLQSNADGKIVSREKMGFHIPCGRFKVARKDSINSVNIFSHVVDCDRVAREV 479
F++RI+W + + ++ E+ I CG+ K+ D+ +FSH++ C+++ R
Sbjct: 31 FKLRITWFEPDPDEQDQVHWVEEE-LPIACGKHKLGITDTTEDRLMFSHLIVCEKIGRCT 207
Query: 480 YKIYPRKGSVWALYGEATLDADGRHFADEGKRCND--IVVFLTNYNEMNGLSMAYLEKVD 537
YK+YPRKG WAL+ + H E R D V L++Y E G+ ++YL K+
Sbjct: 208 YKVYPRKGETWALFKNWDIK---WHMDAESHREYDFEFVEILSDYVEGVGVVVSYLAKLK 378
Query: 538 GYKTVFKRQDTGSHAIR 554
G+ +F R + G+ +
Sbjct: 379 GFVCLFSRMEGGNRTFQ 429
>TC226276 weakly similar to UP|Q6K2F0 (Q6K2F0) Heat shock protein-like,
partial (19%)
Length = 992
Score = 68.9 bits (167), Expect = 6e-12
Identities = 50/136 (36%), Positives = 69/136 (49%)
Frame = +3
Query: 466 FSHVVDCDRVAREVYKIYPRKGSVWALYGEATLDADGRHFADEGKRCNDIVVFLTNYNEM 525
FSH V A IYPRKG VWA+Y + D + ADE D+V L ++ E
Sbjct: 6 FSHKVRWRTGAEGAICIYPRKGDVWAIYRNWSPDWN-ELTADEVIHKFDVVEVLEDFIEG 182
Query: 526 NGLSMAYLEKVDGYKTVFKRQDTGSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLED 585
+G+ + L KV G++TVF IR + ++ M+ SHQIP+ E PE +
Sbjct: 183 HGIDVIPLVKVAGFRTVF-HHHLDPKEIRIIPREEMFRFSHQIPSYVLTG-QEAPEAPKG 356
Query: 586 CWELDPASLPSYLLTI 601
C LDPA+ P LL +
Sbjct: 357 CRVLDPAATPFELLQV 404
>BF424468
Length = 394
Score = 65.9 bits (159), Expect = 5e-11
Identities = 32/78 (41%), Positives = 48/78 (61%)
Frame = +1
Query: 374 DSDFYDFDKDRVERSFKKGQVWAAYDDDDGMPRHYALIDETVSANPFEVRISWLDLQSNA 433
D DF DF++D+ E F Q+WA +D+ D MPR YAL+ + S PF++RI+WL+ S+
Sbjct: 151 DPDFSDFERDKAEDCFAVNQLWAIFDNTDSMPRFYALVKKVYS--PFKLRITWLEPDSDD 324
Query: 434 DGKIVSREKMGFHIPCGR 451
G+I E + CG+
Sbjct: 325 QGEIDWHE-ASLPVACGK 375
>BG507382
Length = 439
Score = 65.9 bits (159), Expect = 5e-11
Identities = 59/172 (34%), Positives = 79/172 (45%), Gaps = 3/172 (1%)
Frame = -1
Query: 82 KKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSALKKGYDAELR---KKEAPTFWT 138
K L LL P+KN A +EA V EA+ VLSD K +D E+ K + +FWT
Sbjct: 439 KTLVRLLDPNKNKLPFA---DEALMRVREAWFVLSDPTRKARFDKEINDAAKTKTTSFWT 269
Query: 139 ACSACRLLHQFERRYVGHSLVCPNCNKSFEAVEAVLSDGSSDEGEKVGVRRNEKREVVLG 198
C C LH++ER+Y +L C NC ++F +S E V E
Sbjct: 268 MCPYCWYLHEYERKYEDCTLRCSNCKRTFHGAAV-----TSPRPEAVAAGNEE--YYCYH 110
Query: 199 VAAPNGVKGKVGGEKGLKGKVGDGDGVGDVGGGGSKKRMRSVGDVFERSKPK 250
V+ P V+ VGGE + + GD G++KRMR V V R K K
Sbjct: 109 VSLP--VRYPVGGE---RCRFGD----------GARKRMR-VKTVANRMKKK 2
>AW184840
Length = 465
Score = 62.0 bits (149), Expect = 7e-10
Identities = 49/130 (37%), Positives = 66/130 (50%), Gaps = 10/130 (7%)
Frame = +1
Query: 292 SRKRVENK-RGLEIGEVRTLKLPIKENAVKSKRGSEVGEERSLKKNVKLAIKEKPEASGK 350
+RK + NK R ++ V K +KEN + SE GE+ S +N ++ ++ E S
Sbjct: 76 ARKEISNKLRQVQSNAVD--KTAMKENGNDFQEVSEKGEKCS--RNSEMCAQDNIEKSED 243
Query: 351 RK---RLELEECGDANGG------ELEVMAVVDSDFYDFDKDRVERSFKKGQVWAAYDDD 401
RK R G E + V+ DF+DF KDR E SF + QVWA YD+D
Sbjct: 244 RKSGSRAIKPFAGSTIAKVSRKFLETTPVDVLYPDFHDFCKDRTEGSFGENQVWAVYDND 423
Query: 402 DGMPRHYALI 411
DGMPR Y LI
Sbjct: 424 DGMPRCYVLI 453
>TC229287 similar to UP|Q8RYF9 (Q8RYF9) Heat shock protein-like, partial (4%)
Length = 817
Score = 60.1 bits (144), Expect = 3e-09
Identities = 55/171 (32%), Positives = 83/171 (48%), Gaps = 11/171 (6%)
Frame = +2
Query: 436 KIVSREKMGFHIPCGRFKV---ARKDSINSVNIFSHVVDC-DRVAREVYKIYPRKGSVWA 491
K V E I GRFK+ AR + + SH V + ++ Y+I+PRKG +WA
Sbjct: 59 KCVKWEDKDMLISIGRFKIKAGARSCTYANTYSVSHQVQVINDGKKKEYEIFPRKGEIWA 238
Query: 492 LYGEATLDADGRHFADEGKRCNDIVVFLTNYNEMNG-----LSMAYLEKVDGYKTVFKRQ 546
LY R++ + KR +D++ + E+ G + + LE V GY +VFKR+
Sbjct: 239 LY---------RNWTTKIKR-SDLLNLEYDIVEVVGEQDLWMDVLPLELVSGYNSVFKRK 388
Query: 547 DT--GSHAIRFLGKDNMWLISHQIPARKSPCIDETPELLEDCWELDPASLP 595
+ A + KD + SHQIPA + +E L WELDP ++P
Sbjct: 389 SNAGSARATKIYWKD-LLRFSHQIPAFE--LTEEQDGNLRGFWELDPGAVP 532
>BM176899 similar to GP|27817868|dbj OJ1081_B12.17 {Oryza sativa (japonica
cultivar-group)}, partial (19%)
Length = 430
Score = 54.7 bits (130), Expect = 1e-07
Identities = 29/71 (40%), Positives = 45/71 (62%)
Frame = +1
Query: 56 ILSATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVL 115
++ T +Y VLGV P A+ ++K Y + +HPDKN AA + F+++GEA++VL
Sbjct: 184 MVKET*YYDVLGVSPTASEAEIKKAYYIKARQVHPDKNPNDPLAA--QNFQVLGEAYQVL 357
Query: 116 SDSALKKGYDA 126
SD A ++ YDA
Sbjct: 358 SDPAQRQAYDA 390
>TC232773 similar to UP|Q9S729 (Q9S729) GlsA, partial (4%)
Length = 420
Score = 54.7 bits (130), Expect = 1e-07
Identities = 41/142 (28%), Positives = 62/142 (42%), Gaps = 18/142 (12%)
Frame = +3
Query: 4 SKGEEEALRLKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVTSLTILSATDWY 63
SKG E+ G+++ + +N+ K R + L E E E+ D Y
Sbjct: 27 SKGVEDNANQLGISDKSINYTNDLKQ-----NRVRLLEMEEEARKEI--------PLDMY 167
Query: 64 TVLGVEPFANSNAVRKQYKKLSLLLHPDKNN------------------THVAAASEEAF 105
+LGVEP + + + Y+K +L HPDK V ++ F
Sbjct: 168 LILGVEPSVSISEINNAYRKAALRHHPDKAGQSLTKNDNGDDQIWKVIAEEVHGDVDQLF 347
Query: 106 KLVGEAFRVLSDSALKKGYDAE 127
K++GEA+ VLSD A + YDAE
Sbjct: 348 KIIGEAYAVLSDPAKRARYDAE 413
>TC231382 weakly similar to GB|AAP21357.1|30102878|BT006549 At1g56300
{Arabidopsis thaliana;} , partial (59%)
Length = 973
Score = 54.3 bits (129), Expect = 2e-07
Identities = 28/73 (38%), Positives = 45/73 (61%), Gaps = 2/73 (2%)
Frame = +3
Query: 58 SATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDK--NNTHVAAASEEAFKLVGEAFRVL 115
S+T +Y VLGV +N + +R+ Y+KL++ HPDK + + ++ F+ + EA+ VL
Sbjct: 132 SSTSYYNVLGVSSDSNVDEIRRAYRKLAMQWHPDKCTRSPSLLGEAKRKFQQIQEAYSVL 311
Query: 116 SDSALKKGYDAEL 128
SDS + YDA L
Sbjct: 312 SDSKKRTMYDAGL 350
>TC220213 similar to UP|Q7XYV0 (Q7XYV0) P58IPK, partial (38%)
Length = 981
Score = 53.9 bits (128), Expect = 2e-07
Identities = 41/128 (32%), Positives = 66/128 (51%), Gaps = 5/128 (3%)
Frame = +1
Query: 3 PSKGEEEALR-LKGLAESKLQTSNNPKSALKYAKRAQRLFPELEGISEMVT----SLTIL 57
P K +EE + L E+KL T + + A++ + A + P+ I E V +L I
Sbjct: 31 PLKIDEELVEALVQRGEAKLLTEDW-EGAVEDLRSAAQKSPQDMNIREAVMRAEKALKIS 207
Query: 58 SATDWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSD 117
D+Y +LG+ A++ +++ YKKL+L HPDK N +E F+ + A+ VLSD
Sbjct: 208 KRKDYYKILGISKTASAADIKRAYKKLALQWHPDK-NVEKREEAEAQFREIAAAYEVLSD 384
Query: 118 SALKKGYD 125
+ YD
Sbjct: 385 EDKRVRYD 408
>TC232105 similar to UP|Q9FEW7 (Q9FEW7) DnaJ like protein, partial (25%)
Length = 315
Score = 52.8 bits (125), Expect = 4e-07
Identities = 27/65 (41%), Positives = 39/65 (59%)
Frame = +1
Query: 61 DWYTVLGVEPFANSNAVRKQYKKLSLLLHPDKNNTHVAAASEEAFKLVGEAFRVLSDSAL 120
D+Y +L V+ A ++K Y+KL++ HPDKN T+ A E FK + EA+ VLSD
Sbjct: 61 DYYKILQVDKHATDEELKKAYRKLAMKWHPDKNPTNKKEA-ETKFKQISEAYEVLSDPQK 237
Query: 121 KKGYD 125
+ YD
Sbjct: 238 RAIYD 252
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.134 0.386
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,675,328
Number of Sequences: 63676
Number of extensions: 305279
Number of successful extensions: 1864
Number of sequences better than 10.0: 182
Number of HSP's better than 10.0 without gapping: 1718
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1803
length of query: 606
length of database: 12,639,632
effective HSP length: 103
effective length of query: 503
effective length of database: 6,081,004
effective search space: 3058745012
effective search space used: 3058745012
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0230.4


