

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0223.9
(324 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
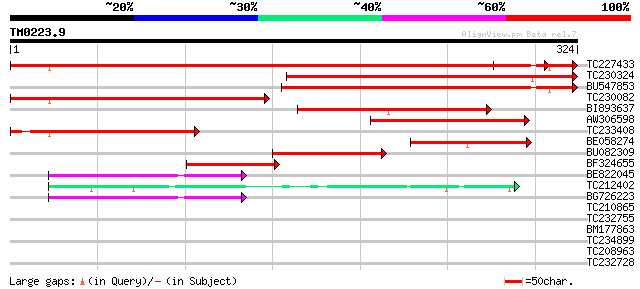
Score E
Sequences producing significant alignments: (bits) Value
TC227433 UP|Q7XZ08 (Q7XZ08) Galactinol synthase, complete 544 e-155
TC230324 homologue to UP|Q7XZ08 (Q7XZ08) Galactinol synthase, pa... 276 9e-75
BU547853 276 1e-74
TC230082 similar to UP|Q9XEJ7 (Q9XEJ7) Galactinol synthase (Frag... 243 9e-65
BI893637 similar to GP|5608499|emb galactinol synthase isoform ... 194 6e-50
AW306598 169 1e-42
TC233408 homologue to UP|Q7XZ08 (Q7XZ08) Galactinol synthase, pa... 164 5e-41
BE058274 similar to GP|20340247|gb| galactinol synthase-like pro... 125 3e-29
BU082309 similar to GP|20340247|gb| galactinol synthase-like pro... 114 5e-26
BF324655 67 8e-12
BE822045 63 2e-10
TC212402 weakly similar to GB|AAO64859.1|29028960|BT005924 At4g3... 62 4e-10
BG726223 60 1e-09
TC210865 30 1.1
TC232755 weakly similar to UP|Q7XAF8 (Q7XAF8) Papillar cell-spec... 29 2.5
BM177863 28 5.6
TC234899 weakly similar to UP|Q9LUJ4 (Q9LUJ4) Gb|AAF26800.1, par... 28 7.4
TC208963 28 7.4
TC232728 27 9.6
>TC227433 UP|Q7XZ08 (Q7XZ08) Galactinol synthase, complete
Length = 1467
Score = 544 bits (1402), Expect = e-155
Identities = 258/330 (78%), Positives = 289/330 (87%), Gaps = 6/330 (1%)
Frame = +1
Query: 1 MAPDLTTAATPITTTEQPLAT---RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVA 57
MAP++TT T IT + +AT RA+VTFLAGNGDYVKGVVGLAKGLRKV+S+YPLVVA
Sbjct: 232 MAPNITTVKTTITDAQAKVATDHGRAYVTFLAGNGDYVKGVVGLAKGLRKVKSMYPLVVA 411
Query: 58 ILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIY 117
+LPDVP++HR IL SQGCIVREI PVYPP+NQTQFA AYYVINYSKLRIWEFVEYSKMIY
Sbjct: 412 VLPDVPQDHRNILTSQGCIVREIEPVYPPENQTQFAMAYYVINYSKLRIWEFVEYSKMIY 591
Query: 118 LDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGP 177
LDGDIQVF+NIDHLFD+PDN FYAV DCFCEP+W HTKQY I YCQQCP KV WP++FGP
Sbjct: 592 LDGDIQVFDNIDHLFDLPDNYFYAVMDCFCEPTWGHTKQYQIGYCQQCPHKVQWPTHFGP 771
Query: 178 RPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVL 237
+PPLYFNAG FVYEPNL TY DLL+T + T PTSFAEQDFLNMYF+DKY+PIPNVYNLVL
Sbjct: 772 KPPLYFNAGMFVYEPNLATYRDLLQTVQVTKPTSFAEQDFLNMYFKDKYRPIPNVYNLVL 951
Query: 238 AMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDW 297
AMLWRHPENV+L+KVKVVHYCA GSKPWR+TG+EENM+R+DIKMLVKKWWDIYEDETLD+
Sbjct: 952 AMLWRHPENVELEKVKVVHYCAAGSKPWRYTGKEENMEREDIKMLVKKWWDIYEDETLDY 1131
Query: 298 NYINPSYTEK---VLIDGGGVKNVSAPSAA 324
N NP +K L++ G VK V APSAA
Sbjct: 1132N--NPLNVDKFTAALMEVGEVKFVRAPSAA 1215
Score = 42.7 bits (99), Expect = 2e-04
Identities = 21/32 (65%), Positives = 23/32 (71%)
Frame = +1
Query: 277 KDIKMLVKKWWDIYEDETLDWNYINPSYTEKV 308
+DIKMLVKK WDIYEDETLD N + KV
Sbjct: 1 EDIKMLVKKRWDIYEDETLDNNKPTKIFLTKV 96
>TC230324 homologue to UP|Q7XZ08 (Q7XZ08) Galactinol synthase, partial (43%)
Length = 748
Score = 276 bits (706), Expect = 9e-75
Identities = 128/168 (76%), Positives = 141/168 (83%), Gaps = 2/168 (1%)
Frame = +2
Query: 159 IKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQDFL 218
I YCQQCPDKV WPS+FG +PPLYFNAG F YEPNL TY LL+T + PTSFAEQDFL
Sbjct: 2 IGYCQQCPDKVQWPSHFGTKPPLYFNAGMFDYEPNLNTYRHLLQTVQVIKPTSFAEQDFL 181
Query: 219 NMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMDRKD 278
NMYF+DKYKPIPNVYNLVLAMLWRHPENV+LDKV+VVHYCA GSKPWRFTG+EENMDR+D
Sbjct: 182 NMYFKDKYKPIPNVYNLVLAMLWRHPENVELDKVQVVHYCAAGSKPWRFTGKEENMDRED 361
Query: 279 IKMLVKKWWDIYEDETLDW--NYINPSYTEKVLIDGGGVKNVSAPSAA 324
IKMLVKKWWDIYEDETLD+ N +N L+D GG + V APSAA
Sbjct: 362 IKMLVKKWWDIYEDETLDYNNNSVNVERFTSALLDAGGFQFVPAPSAA 505
>BU547853
Length = 686
Score = 276 bits (705), Expect = 1e-74
Identities = 128/172 (74%), Positives = 148/172 (85%), Gaps = 3/172 (1%)
Frame = -2
Query: 156 QYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQ 215
QY I YCQQCP KV WP+++GP+PPLYFNAG FVYEPNL+TY DLL+T + T PTSFAEQ
Sbjct: 685 QYQIGYCQQCPHKVQWPTHYGPKPPLYFNAGMFVYEPNLDTYRDLLQTVQVTKPTSFAEQ 506
Query: 216 DFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKVVHYCAFGSKPWRFTGEEENMD 275
DFLNMYF+DKY+PIPNVYNLVLAMLWRHPENV+L+KVKVVHYCA GSKP R+TG+EENM+
Sbjct: 505 DFLNMYFKDKYRPIPNVYNLVLAMLWRHPENVELEKVKVVHYCAAGSKPXRYTGKEENME 326
Query: 276 RKDIKMLVKKWWDIYEDETLDWNYINPSYTEK---VLIDGGGVKNVSAPSAA 324
R+DIKMLVKKWWDIYEDETLD+N NP ++ L++ G VK V APSAA
Sbjct: 325 REDIKMLVKKWWDIYEDETLDYN--NPFNVDRFTAALLEVGEVKFVRAPSAA 176
>TC230082 similar to UP|Q9XEJ7 (Q9XEJ7) Galactinol synthase (Fragment),
partial (39%)
Length = 647
Score = 243 bits (620), Expect = 9e-65
Identities = 119/153 (77%), Positives = 129/153 (83%), Gaps = 5/153 (3%)
Frame = +2
Query: 1 MAPDLTTAATPITTTEQPLAT-----RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLV 55
M P++TT +TT + P A RAFVTFLAGNGDYVKGVVGLAKGLRK +S+YPLV
Sbjct: 107 MPPNITTVVANVTTEQLPKAHGGSSGRAFVTFLAGNGDYVKGVVGLAKGLRKAKSMYPLV 286
Query: 56 VAILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKM 115
VA+LPDVPEEHR+IL SQGCIVREI PVYPP+NQTQFA AYYVINYSKLRIWEFVEY K
Sbjct: 287 VAVLPDVPEEHREILKSQGCIVREIEPVYPPENQTQFAMAYYVINYSKLRIWEFVEYKKT 466
Query: 116 IYLDGDIQVFENIDHLFDMPDNQFYAVKDCFCE 148
IYLDGDIQVF NIDHL D+PDN FYAV DCFCE
Sbjct: 467 IYLDGDIQVFGNIDHLVDLPDNYFYAVMDCFCE 565
>BI893637 similar to GP|5608499|emb galactinol synthase isoform GolS-2
{Ajuga reptans}, partial (36%)
Length = 425
Score = 194 bits (492), Expect = 6e-50
Identities = 91/141 (64%), Positives = 102/141 (71%), Gaps = 30/141 (21%)
Frame = +3
Query: 165 CPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPTSFAEQ--------- 215
CP KV WP++FGP+PPLYFNAG FVYEPNL TY DLL+T + T PTSFAEQ
Sbjct: 3 CPHKVQWPTHFGPKPPLYFNAGIFVYEPNLATYRDLLQTVQVTQPTSFAEQVFNY*VN*S 182
Query: 216 ---------------------DFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKV 254
DFLNMYF+DKY+PIPNVYNLVLAMLWRHPENV+LDKVKV
Sbjct: 183 YTILIYYKV*RVYLCWIIGLQDFLNMYFKDKYRPIPNVYNLVLAMLWRHPENVELDKVKV 362
Query: 255 VHYCAFGSKPWRFTGEEENMD 275
VHYCA GSKPWR+TG+EENM+
Sbjct: 363 VHYCAAGSKPWRYTGKEENME 425
>AW306598
Length = 388
Score = 169 bits (429), Expect = 1e-42
Identities = 76/91 (83%), Positives = 84/91 (91%)
Frame = +1
Query: 207 TTPTSFAEQDFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQLDKVKVVHYCAFGSKPWR 266
TTPTSFAEQDFLNMYF+D YKPIP YNLVLAMLWRHPENV+LD+VKVVHYCA GSKPWR
Sbjct: 1 TTPTSFAEQDFLNMYFKDIYKPIPLNYNLVLAMLWRHPENVKLDQVKVVHYCAAGSKPWR 180
Query: 267 FTGEEENMDRKDIKMLVKKWWDIYEDETLDW 297
+TG+EENM R+DIKMLVKKWWDIY D +LD+
Sbjct: 181 YTGKEENMQREDIKMLVKKWWDIYNDASLDY 273
>TC233408 homologue to UP|Q7XZ08 (Q7XZ08) Galactinol synthase, partial (33%)
Length = 527
Score = 164 bits (415), Expect = 5e-41
Identities = 84/111 (75%), Positives = 93/111 (83%), Gaps = 3/111 (2%)
Frame = +2
Query: 1 MAPDLTTAATPITTTEQPLAT---RAFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVA 57
MAP++TT +T + A RA+VTFLAGNGDYVKGVVGLAKGLRKV+S+YPLVVA
Sbjct: 206 MAPNITT----VTDAQAKAAGGRGRAYVTFLAGNGDYVKGVVGLAKGLRKVKSMYPLVVA 373
Query: 58 ILPDVPEEHRKILLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWE 108
+LPDVPE HR IL SQGCIVREI PVYPP+NQTQFA AYYVINYSKLRIWE
Sbjct: 374 VLPDVPEHHRNILTSQGCIVREIEPVYPPENQTQFAMAYYVINYSKLRIWE 526
>BE058274 similar to GP|20340247|gb| galactinol synthase-like protein
{Thellungiella halophila}, partial (20%)
Length = 327
Score = 125 bits (313), Expect = 3e-29
Identities = 61/100 (61%), Positives = 68/100 (68%), Gaps = 31/100 (31%)
Frame = +3
Query: 230 PNVYNLVLAMLWRHPENVQLDKVKVVHYCAF----------------------------- 260
PNVYNLVLAMLWRHPENV+L+KVKVVHYCA
Sbjct: 6 PNVYNLVLAMLWRHPENVELEKVKVVHYCAAVSVQF*LK*NWNLNFGIQ*ERNFNLVELV 185
Query: 261 --GSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDETLDWN 298
GSKPWR+TG+EENM+R+DIKMLVKKWWDIYEDETLD+N
Sbjct: 186 GQGSKPWRYTGKEENMEREDIKMLVKKWWDIYEDETLDYN 305
>BU082309 similar to GP|20340247|gb| galactinol synthase-like protein
{Thellungiella halophila}, partial (18%)
Length = 421
Score = 114 bits (286), Expect = 5e-26
Identities = 48/65 (73%), Positives = 54/65 (82%)
Frame = +3
Query: 151 WRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLETYHDLLKTCEATTPT 210
W HT QY I YCQQCP KV WP++FGP+PPLYFNAG FVYEPNL+TY DLL+T + T PT
Sbjct: 3 WGHTLQYQIGYCQQCPHKVQWPTHFGPKPPLYFNAGMFVYEPNLDTYRDLLQTVQVTKPT 182
Query: 211 SFAEQ 215
SFAEQ
Sbjct: 183 SFAEQ 197
>BF324655
Length = 160
Score = 67.4 bits (163), Expect = 8e-12
Identities = 31/53 (58%), Positives = 40/53 (74%)
Frame = +2
Query: 102 SKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMPDNQFYAVKDCFCEPSWRHT 154
SKL I E VE++ MIYLDGDIQV++NIDH D+PD+ YA +DC CE + H+
Sbjct: 2 SKLLICE*VEFAMMIYLDGDIQVYDNIDHWCDLPDDC*YAARDCICEATSGHS 160
>BE822045
Length = 632
Score = 63.2 bits (152), Expect = 2e-10
Identities = 37/113 (32%), Positives = 58/113 (50%)
Frame = +1
Query: 23 AFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKILLSQGCIVREIVP 82
A+ T L + YV G + LA+ L + + L++ I + R+ L G +R I
Sbjct: 202 AYATVLHSSEAYVCGAITLAQSLLQTGTKRDLILLIDKFISVRKREALSEAGWKIRIITR 381
Query: 83 VYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMP 135
+ PK + + Y NYSK R+W+ +Y K+I++D DI V N+D LF P
Sbjct: 382 IRNPKAEKGSYNEY---NYSKFRLWQLTDYDKVIFIDSDIIVLRNLDILFHXP 531
>TC212402 weakly similar to GB|AAO64859.1|29028960|BT005924 At4g33330
{Arabidopsis thaliana;} , partial (37%)
Length = 1146
Score = 62.0 bits (149), Expect = 4e-10
Identities = 69/291 (23%), Positives = 117/291 (39%), Gaps = 22/291 (7%)
Frame = +2
Query: 23 AFVTFLAGNGDYVKGVVGLAKGL-----------RKVQSIYPLVVAILPDVPEEHRKI-- 69
A+VT L + YV G + LA+ + + L + +L D ++ I
Sbjct: 2 AYVTVLHSSEAYVCGAIALAQSILQHNNNNNNNNNNNNNYTKLDLLLLADESIGYKSIRG 181
Query: 70 LLSQGCIVREIVPVYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENID 129
L + G ++ I + P Q +Y NYS+LRIW+ Y K+I+LD D+ V ++ID
Sbjct: 182 LKAAGWKIKRIKRILNPYAQKG---SYNEWNYSRLRIWQLTMYDKIIFLDADLLVLKSID 352
Query: 130 HLFDMPDNQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFV 189
LF P Q S P++F F +G V
Sbjct: 353 GLFAYP--------------------QLSAS-----------PNDFS-----LFKSGLMV 424
Query: 190 YEPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQ- 248
EP+ + DL+K + +Q +N F ++ +P N + + R +V+
Sbjct: 425 IEPSTCMFEDLMKKSLEVKSYNGGDQGLVNEVFTWWHR-LPTKVNYLKSFEEREGNDVKE 601
Query: 249 --LDKVKVVHYCAFGSKPWR-FTGEEENMDRKDIKMLVK-----KWWDIYE 291
+ + V+HY G KPW + + N D ++ + WW +Y+
Sbjct: 602 EIPEDLYVMHY--LGLKPWMCYRDYDCNWDMNELHVFASDLAHHMWWQVYD 748
>BG726223
Length = 412
Score = 60.5 bits (145), Expect = 1e-09
Identities = 34/113 (30%), Positives = 58/113 (51%)
Frame = +3
Query: 23 AFVTFLAGNGDYVKGVVGLAKGLRKVQSIYPLVVAILPDVPEEHRKILLSQGCIVREIVP 82
A+ T L + YV G + LA+ L + + L++ + + R+ L G +R I
Sbjct: 21 AYATVLHSSEGYVCGAITLAQTLLQTGTKRDLILLLDTSISVAKRRSLELSGWKIRLITR 200
Query: 83 VYPPKNQTQFAHAYYVINYSKLRIWEFVEYSKMIYLDGDIQVFENIDHLFDMP 135
+ P+ + + Y NYSK R+W+ +Y ++I++D DI V N+D LF P
Sbjct: 201 IRNPRAENGTYNEY---NYSKFRLWQLTDYERVIFIDADIIVLRNLDILFHFP 350
>TC210865
Length = 1111
Score = 30.4 bits (67), Expect = 1.1
Identities = 16/36 (44%), Positives = 21/36 (57%), Gaps = 6/36 (16%)
Frame = -2
Query: 153 HTKQYSIKYCQ-----QCPDKVDWPS-NFGPRPPLY 182
H + I +CQ QCPD++D PS NF P P L+
Sbjct: 558 HRSPFQILHCQERKKEQCPDQLDPPS*NFQPTPYLW 451
>TC232755 weakly similar to UP|Q7XAF8 (Q7XAF8) Papillar cell-specific pectin
methylesterase-like protein, partial (7%)
Length = 718
Score = 29.3 bits (64), Expect = 2.5
Identities = 36/132 (27%), Positives = 53/132 (39%), Gaps = 14/132 (10%)
Frame = +1
Query: 137 NQFYAVKDCFCEPSWRHTKQYSIKYCQQCPDKVDWPSNFGPRPPLYFNAGFFVYEPNLE- 195
NQ+ ++ D +C S R + S K+ + PS++ F+ E NLE
Sbjct: 241 NQYGSIYD-YCRISVRKSLSQSRKFLNNMYSYLQNPSSYSQSTIRALEDCQFLAELNLEY 417
Query: 196 --TYHDLLKTCEATTPTSFAE--QDFLNMYFRDKY---------KPIPNVYNLVLAMLWR 242
T HD + A PTS AE L+ ++ P P V N + L
Sbjct: 418 LSTTHDTVDKASAVLPTSQAEDVHTLLSAVLTNQQTCLDGLQTSAPDPRVKNDLSLQL-- 591
Query: 243 HPENVQLDKVKV 254
ENV+LD V +
Sbjct: 592 -AENVKLDSVSL 624
>BM177863
Length = 422
Score = 28.1 bits (61), Expect = 5.6
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = +1
Query: 299 YINPSYTEKVLIDGGGVKNVSAPSA 323
Y+NP+ E++ I GGVK + P+A
Sbjct: 136 YVNPTSAEEIRIGLGGVKPIMTPAA 210
>TC234899 weakly similar to UP|Q9LUJ4 (Q9LUJ4) Gb|AAF26800.1, partial (36%)
Length = 767
Score = 27.7 bits (60), Expect = 7.4
Identities = 24/107 (22%), Positives = 47/107 (43%), Gaps = 4/107 (3%)
Frame = +2
Query: 191 EPNLETYHDLLKTCEATTPTSFAEQDFLNMYFRDKYKPIPNVYNLVLAMLWRHPENVQ-- 248
+PN+ TYH LLK C + L+ F++ P Y+L++ L + +
Sbjct: 350 KPNVGTYHPLLKMC-CKKKRMKVLKFLLDHMFKNDISPDLATYSLLVNALCKTGKVADAY 526
Query: 249 --LDKVKVVHYCAFGSKPWRFTGEEENMDRKDIKMLVKKWWDIYEDE 293
L+++ + + S GE E++ + K V++W D + +
Sbjct: 527 SFLEEMVLKGFTPKPSTLKGLAGELESLSMLEEKERVEEWMDRFSQK 667
>TC208963
Length = 1163
Score = 27.7 bits (60), Expect = 7.4
Identities = 13/45 (28%), Positives = 21/45 (45%), Gaps = 12/45 (26%)
Frame = -1
Query: 75 CIVREIVPVYPPKNQTQFAHA------------YYVINYSKLRIW 107
C+ ++ PK T+F HA +++INYSK +W
Sbjct: 521 CVASTLLVAVLPKCITRFVHAVTNFSLCGISHVFFLINYSKYNVW 387
>TC232728
Length = 807
Score = 27.3 bits (59), Expect = 9.6
Identities = 16/36 (44%), Positives = 23/36 (63%), Gaps = 3/36 (8%)
Frame = +2
Query: 222 FRDKYKPIPNVYNLV---LAMLWRHPENVQLDKVKV 254
FRDK PN NLV L++L R E+++L+ VK+
Sbjct: 329 FRDKVNSSPNELNLVSNELSILRRENEDLKLEIVKL 436
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,438,664
Number of Sequences: 63676
Number of extensions: 260644
Number of successful extensions: 1232
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1224
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1231
length of query: 324
length of database: 12,639,632
effective HSP length: 97
effective length of query: 227
effective length of database: 6,463,060
effective search space: 1467114620
effective search space used: 1467114620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Lotus: description of TM0223.9


