

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
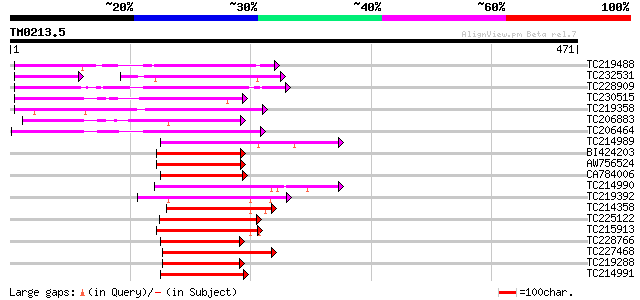
Query= TM0213.5
(471 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219488 UP|Q8S900 (Q8S900) Syringolide-induced protein 1-3-1A, ... 139 2e-33
TC232531 UP|Q8S8Z9 (Q8S8Z9) Syringolide-induced protein 1-3-1B, ... 108 8e-33
TC228909 similar to GB|AAS09999.1|41618996|AY519529 MYB transcri... 122 3e-28
TC230515 similar to GB|AAS09999.1|41618996|AY519529 MYB transcri... 121 6e-28
TC219358 homologue to PIR|F96527|F96527 protein F27J15.20 [impor... 120 1e-27
TC206883 similar to GB|AAS10003.1|41619012|AY519533 MYB transcri... 119 3e-27
TC206464 similar to GB|AAS58519.1|45357120|AY550308 MYB transcri... 118 6e-27
TC214989 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1... 99 5e-21
BI424203 98 7e-21
AW756524 97 2e-20
CA784006 96 3e-20
TC214990 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1... 96 3e-20
TC219392 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1... 96 5e-20
TC214358 homologue to UP|Q84UB0 (Q84UB0) Transcription factor My... 95 6e-20
TC225122 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1... 93 3e-19
TC215913 similar to GB|AAS09986.1|41618940|AY519516 MYB transcri... 91 9e-19
TC228766 similar to GB|AAS99672.1|46518381|BT012528 At5g56840 {A... 89 3e-18
TC227468 similar to UP|Q8H1D0 (Q8H1D0) Transcription factor MYBS... 89 3e-18
TC219288 similar to GB|AAS99672.1|46518381|BT012528 At5g56840 {A... 88 9e-18
TC214991 similar to GB|AAS09980.1|41618916|AY519510 MYB transcri... 87 1e-17
>TC219488 UP|Q8S900 (Q8S900) Syringolide-induced protein 1-3-1A, complete
Length = 1338
Score = 139 bits (351), Expect = 2e-33
Identities = 88/223 (39%), Positives = 118/223 (52%), Gaps = 3/223 (1%)
Frame = +2
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEG--PA 61
WT D+ E AL VP+D PDRWE+IA V G+ E+ E Y LV V I+ G
Sbjct: 257 WTRYHDKLFERALLVVPEDLPDRWEKIADQVPGKSAVEVREHYEALVHDVFEIDSGRVEV 436
Query: 62 PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKK 121
P ++ D V P PS I+ + N S K+ K+
Sbjct: 437 PSYVDDSVAMP----PSGGAGISTWDNAN--------------------QISFGSKL-KQ 541
Query: 122 ISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLN 181
N+ +K WT++EHR FL GL K+GKG W++ISR + ++TPTQ+ASHAQKY+LR N
Sbjct: 542 QGENERKKGTPWTEEEHRLFLIGLSKFGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQN 721
Query: 182 STPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQN 224
S K RKR+SIHD+T D++ P + Q P SPQQ+
Sbjct: 722 SVKKERKRSSIHDITTVDSNSAPM--PIDQTWVPPPGGSPQQS 844
>TC232531 UP|Q8S8Z9 (Q8S8Z9) Syringolide-induced protein 1-3-1B, partial
(98%)
Length = 1225
Score = 108 bits (270), Expect(2) = 8e-33
Identities = 62/154 (40%), Positives = 86/154 (55%), Gaps = 17/154 (11%)
Frame = +2
Query: 93 EGPAPDFIVDRVPEPVPNDPSSSQKIT--------------KKISPNQNEKRVLWTDDEH 138
E P ++ D V P PS +I+ K+ N+ +K WT++EH
Sbjct: 407 EFKVPSYVDDSVATP----PSGGAEISTWDNANQISFGSKPKQQGDNERKKGTPWTEEEH 574
Query: 139 RNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTID 198
R FL GL K+GKG W++ISR + ++TPTQ+ASHAQKY+LR NS K RKR+SIHD+T
Sbjct: 575 RLFLIGLSKFGKGDWRSISRNVVVTRTPTQVASHAQKYFLRQNSVKKERKRSSIHDITTV 754
Query: 199 DTDLVP---QQNQVPQLNAPMQELSPQQNGVQLD 229
D++ VP QN VP +Q+ + LD
Sbjct: 755 DSNSVPVPIDQNWVPPPGGSVQQSLQYPSSNMLD 856
Score = 50.4 bits (119), Expect(2) = 8e-33
Identities = 26/58 (44%), Positives = 33/58 (56%), Gaps = 1/58 (1%)
Frame = +3
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPA 61
WT D+ E AL VP+D PDRWE+IA V G+ E+ E Y LV V I+ P+
Sbjct: 237 WTRYHDKLFERALLVVPEDLPDRWEKIADQVPGKSAVEVREHYEALVHDVFEIDSRPS 410
>TC228909 similar to GB|AAS09999.1|41618996|AY519529 MYB transcription factor
{Arabidopsis thaliana;} , partial (60%)
Length = 1612
Score = 122 bits (307), Expect = 3e-28
Identities = 80/230 (34%), Positives = 123/230 (52%), Gaps = 1/230 (0%)
Frame = +1
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
WTP +++ E ALA D PDRW ++AA + G+ +++++YR L V IE G
Sbjct: 559 WTPQENKLFENALAVFDKDTPDRWLKVAALIPGKTVGDVIKQYRELEEDVSVIESG---- 726
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
FI P+P +++ T + N G F + + + + S
Sbjct: 727 FI------PLPGY-TATDSFTLEWVNNQGFGGLRQFY----------GVTGKRGASTRPS 855
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
+ +K V WT +EHR FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R S
Sbjct: 856 EQERKKGVPWTKEEHRQFLMGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQLSG 1035
Query: 184 PKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSPQQNGVQLDAPMQ 233
K +KR+SIHD+T+ + +P+ P + + SP + ++ P Q
Sbjct: 1036GKDKKRSSIHDITMVN---LPEAKS-PSSESNNRPYSPDHSVKDVNPPSQ 1173
>TC230515 similar to GB|AAS09999.1|41618996|AY519529 MYB transcription factor
{Arabidopsis thaliana;} , partial (50%)
Length = 946
Score = 121 bits (304), Expect = 6e-28
Identities = 76/196 (38%), Positives = 105/196 (52%), Gaps = 3/196 (1%)
Frame = +1
Query: 5 WTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPD 63
WT +++ E ALA D PDRW ++AA + G+ +++++YR L V IE G P
Sbjct: 163 WTREENKKFESALAIYDKDTPDRWLRVAAMLPGKTVYDVIKQYRELEEDVCEIEAGRIP- 339
Query: 64 FIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKIS 123
VP P+SS T K+ N D + T + S
Sbjct: 340 ---------VPGYPTSS--FTLKMVDN-----------------QCYDACRKKPATLRSS 435
Query: 124 PNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLR--LN 181
+ +K V WT++EHR FL GL KYGKG W+ ISR F+ +KTPTQ+ASHAQKYY+R L+
Sbjct: 436 DQERKKGVPWTEEEHRRFLMGLLKYGKGDWRNISRNFVVTKTPTQVASHAQKYYIRQKLS 615
Query: 182 STPKRRKRASIHDLTI 197
++R SIHD+TI
Sbjct: 616 GGKDNKRRPSIHDITI 663
>TC219358 homologue to PIR|F96527|F96527 protein F27J15.20 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(39%)
Length = 1195
Score = 120 bits (301), Expect = 1e-27
Identities = 74/217 (34%), Positives = 123/217 (56%), Gaps = 7/217 (3%)
Frame = +3
Query: 5 WTPAQDRALELALAT--VPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPA 61
W+ +++A E A+A + + + ++WE+IA+AV + E+ + Y++LV V IE G
Sbjct: 51 WSYDEEKAFENAIAMHWIEESSKEQWEKIASAVPSKSMEEVKQHYQVLVEDVSAIEAGHI 230
Query: 62 --PDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFI-VDRVPEPVPNDPSSSQKI 118
P++ D + SS+ + N G + +D SS +
Sbjct: 231 SFPNYASDETTSSNKDFHGSSKATSSDKRSNCNYGSGFSGLGLDSTTH------SSGKGG 392
Query: 119 TKKISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYL 178
+ S + K + WT++EHR FL GL K+GKG W++ISR F+ S+TPTQ+ASHAQKY++
Sbjct: 393 LSRSSEQERRKGIPWTEEEHRLFLLGLDKFGKGDWRSISRNFVISRTPTQVASHAQKYFI 572
Query: 179 RLNSTPKRRKRASIHDLT-IDDTDLVPQQNQVPQLNA 214
RLNS + R+R+SIHD+T +++ D+ Q + L++
Sbjct: 573 RLNSMNRDRRRSSIHDITSVNNGDVASSQAPITGLHS 683
>TC206883 similar to GB|AAS10003.1|41619012|AY519533 MYB transcription factor
{Arabidopsis thaliana;} , partial (51%)
Length = 1224
Score = 119 bits (298), Expect = 3e-27
Identities = 73/189 (38%), Positives = 106/189 (55%), Gaps = 3/189 (1%)
Frame = +3
Query: 11 RALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGPAPDFIVDRV 69
+ E ALA D PDRW ++A + G+ +++ +Y+ L + V NIE G P
Sbjct: 3 KLFENALAVHDKDTPDRWHKVAEMIPGKTVVDVIRQYKELEVDVSNIEAGLIP------- 161
Query: 70 PEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITKKISPNQNEK 129
VP S++ ISP F +D V P + K + P ++E+
Sbjct: 162 ---VPGYSSTA------ISP---------FTLDWVNTPGYDGFKGCGKRPSSVRPIEHER 287
Query: 130 R--VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRR 187
+ V WT++EH+ FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R S K +
Sbjct: 288 KKGVPWTEEEHKLFLLGLKKYGKGDWRNISRNFVITRTPTQVASHAQKYFIRQLSGGKDK 467
Query: 188 KRASIHDLT 196
+RASIHD+T
Sbjct: 468 RRASIHDIT 494
>TC206464 similar to GB|AAS58519.1|45357120|AY550308 MYB transcription factor
{Arabidopsis thaliana;} , partial (61%)
Length = 1156
Score = 118 bits (295), Expect = 6e-27
Identities = 77/214 (35%), Positives = 108/214 (49%), Gaps = 3/214 (1%)
Frame = +3
Query: 2 SDSWTPAQDRALELALATVPDDAPDRWEQIAAAV-GRLPAEIMERYRLLVLHVDNIEEGP 60
S WT ++ E ALA +D PDRW ++AA + G+ +++++YR L V IE G
Sbjct: 591 STEWTREDNKKFESALAIYDNDTPDRWFKVAAMIPGKTVFDVIKQYRELEEDVSEIEAGR 770
Query: 61 APDFIVDRVPEPVPNDPSSSQKITKKISPNIEEGPAPDFIVDRVPEPVPNDPSSSQKITK 120
P +P +SS + N D +
Sbjct: 771 VP----------IPGYLASSFTFELVDNHNY-------------------DGCRRRLAPV 863
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
+ S + +K V WT+DEHR FL GL KYGKG W+ ISR F+ +KTPTQ+ASHAQKYY+R
Sbjct: 864 RGSDQERKKGVPWTEDEHRRFLMGLLKYGKGDWRNISRNFVVTKTPTQVASHAQKYYIRQ 1043
Query: 181 N-STPKRRKRASIHDL-TIDDTDLVPQQNQVPQL 212
S K ++R SIHD+ T++ T+ PQL
Sbjct: 1044KVSGGKDKRRPSIHDITTVNLTETSASDKNKPQL 1145
>TC214989 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(42%)
Length = 1511
Score = 98.6 bits (244), Expect = 5e-21
Identities = 55/163 (33%), Positives = 87/163 (52%), Gaps = 11/163 (6%)
Frame = +3
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ ++ V WT++EH+ FL GL+K GKG W+ IS+ ++ ++TPTQ+ASHAQKY+LR ++ +
Sbjct: 339 ERKRGVPWTEEEHKLFLVGLQKVGKGDWRGISKNYVKTRTPTQVASHAQKYFLRRSNLNR 518
Query: 186 RRKRASIHDLTIDDTDLVPQ-----QNQVPQLNAPMQELSPQQNGVQLDAPMQQV----- 235
RR+R+S+ D+T D +P QNQ ++ Q + P + P V
Sbjct: 519 RRRRSSLFDITTDTVSAIPMEEEQVQNQDTLCHSQQQPVFPAETSKINGFPAMPVYQFGV 698
Query: 236 -SSQQNQVDALDFPMQQLSFQQNGDPPLNFPHQVFPQQNCATP 277
SS V PM++L+ Q N P++ TP
Sbjct: 699 GSSGVISVQGTGNPMEELTLGQGNVEKHNVPNKASTVSGIITP 827
>BI424203
Length = 411
Score = 98.2 bits (243), Expect = 7e-21
Identities = 44/74 (59%), Positives = 58/74 (77%)
Frame = +3
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
S + K + WT+DEHR FL GL KYGKG ++ISR F+ ++TPTQ+ASHAQKY++RLNS
Sbjct: 45 SDQERRKGIAWTEDEHRLFLLGLDKYGKGDRRSISRNFVVTRTPTQVASHAQKYFIRLNS 224
Query: 183 TPKRRKRASIHDLT 196
K R+R+SIHD+T
Sbjct: 225 MNKDRRRSSIHDIT 266
>AW756524
Length = 256
Score = 97.1 bits (240), Expect = 2e-20
Identities = 44/74 (59%), Positives = 57/74 (76%)
Frame = +2
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
S + K + WT+DEHR L GL+KYGKG W++ISR F+ + TPTQ+ASHAQKY++RLNS
Sbjct: 26 SDQERLKGIAWTEDEHRLLLLGLEKYGKGDWRSISRNFVVT*TPTQVASHAQKYFIRLNS 205
Query: 183 TPKRRKRASIHDLT 196
K R R+SIHD+T
Sbjct: 206 MNKDRMRSSIHDVT 247
>CA784006
Length = 409
Score = 96.3 bits (238), Expect = 3e-20
Identities = 41/72 (56%), Positives = 58/72 (79%)
Frame = +3
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ +K V WT++EHR FL GLKKYGKG W+ ISR F+ ++TPTQ+ASHAQKY++R + K
Sbjct: 90 ERKKGVPWTEEEHRQFLLGLKKYGKGDWRNISRNFVTTRTPTQVASHAQKYFIRQLTGGK 269
Query: 186 RRKRASIHDLTI 197
++R+SIHD+T+
Sbjct: 270 DKRRSSIHDITM 305
>TC214990 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(59%)
Length = 1443
Score = 96.3 bits (238), Expect = 3e-20
Identities = 55/171 (32%), Positives = 96/171 (55%), Gaps = 14/171 (8%)
Frame = +3
Query: 121 KISPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRL 180
++ + ++ V WT++EH+ FL GL+K GKG W+ IS+ ++ ++TPTQ+ASHAQKY+LR
Sbjct: 330 RLRERERKRGVPWTEEEHKLFLVGLQKVGKGDWRGISKNYVKTRTPTQVASHAQKYFLRR 509
Query: 181 NSTPKRRKRASIHDLTIDDTDLVPQQNQVPQLNAPM---QELSP-------QQNGVQLDA 230
++ +RR+R+S+ D+T D +P + + Q + Q+ SP + NG +
Sbjct: 510 SNLNRRRRRSSLFDITTDTVSAIPMEGEQVQNQDTLSHSQQQSPLFPAETSKINGFPM-M 686
Query: 231 PMQQVSSQQNQVDALD----FPMQQLSFQQNGDPPLNFPHQVFPQQNCATP 277
P+ Q + V ++ PM++L+ Q N P++V + TP
Sbjct: 687 PVYQFGFGSSGVISVQGGNGNPMEELTLGQGNVEKHNVPNKVSTVSDIITP 839
>TC219392 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(28%)
Length = 918
Score = 95.5 bits (236), Expect = 5e-20
Identities = 52/147 (35%), Positives = 82/147 (55%), Gaps = 19/147 (12%)
Frame = +3
Query: 107 PVPNDPSSSQKITKKISPNQNEKR----VLWTDDEHRNFLRGLKKYGKGQWQTISREFLP 162
P P+DP++ + P+++ + V WT++EHR FL GL+ GKG W+ ISR F+
Sbjct: 303 PPPHDPNAGYASDDVVHPSRHTRERKRGVPWTEEEHRLFLLGLQNIGKGDWRGISRNFVK 482
Query: 163 SKTPTQIASHAQKYYLRLNSTPKRRKRASIHDLTID-----------DTDLVPQQNQVPQ 211
++TPTQ+ASHAQKY+LR ++ +RR+R+S+ D+T D + ++P P
Sbjct: 483 TRTPTQVASHAQKYFLRRHTQNRRRRRSSLFDITTDSVMEPWPEKEEEQVVLPSARSKPV 662
Query: 212 LNAP----MQELSPQQNGVQLDAPMQQ 234
L P M EL + L P+ +
Sbjct: 663 LPVPPSSKMAELDLSGKSLSLKLPVSK 743
>TC214358 homologue to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(24%)
Length = 940
Score = 95.1 bits (235), Expect = 6e-20
Identities = 46/96 (47%), Positives = 68/96 (69%), Gaps = 5/96 (5%)
Frame = +2
Query: 131 VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRRKRA 190
V WT++EHR FL GL K GKG W+ ISR F+ ++TPTQ+ASHAQKY+LR ++ +RR+R+
Sbjct: 5 VPWTEEEHRLFLLGLHKVGKGDWRGISRNFVKTRTPTQVASHAQKYFLRRHNQNRRRRRS 184
Query: 191 SIHDLTID---DTDLVPQQNQVPQ--LNAPMQELSP 221
S+ D+T D ++ + ++ QVPQ + AP+ P
Sbjct: 185 SLFDITTDTVMESSTIMEEEQVPQETMAAPIPAAYP 292
>TC225122 similar to UP|Q84UB0 (Q84UB0) Transcription factor Myb1, partial
(40%)
Length = 1301
Score = 92.8 bits (229), Expect = 3e-19
Identities = 41/86 (47%), Positives = 62/86 (71%), Gaps = 1/86 (1%)
Frame = +1
Query: 125 NQNEKR-VLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNST 183
N++ KR + WT++EH+ FL GL+K GKG W+ ISR ++ ++TPTQ+ASHAQKY+LR +
Sbjct: 286 NRDRKRGIPWTEEEHKLFLVGLQKVGKGDWRGISRNYVKTRTPTQVASHAQKYFLRRTNL 465
Query: 184 PKRRKRASIHDLTIDDTDLVPQQNQV 209
+RR+R+S+ D+T D P + V
Sbjct: 466 NRRRRRSSLFDITTDSVSTTPMEEGV 543
>TC215913 similar to GB|AAS09986.1|41618940|AY519516 MYB transcription factor
{Arabidopsis thaliana;} , partial (66%)
Length = 1545
Score = 91.3 bits (225), Expect = 9e-19
Identities = 43/95 (45%), Positives = 67/95 (70%), Gaps = 7/95 (7%)
Frame = +2
Query: 123 SPNQNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNS 182
S + +K V WT++EHR FL GL+K GKG W+ I+R ++ S+TPTQ+ASHAQKY++R ++
Sbjct: 479 SSRERKKGVPWTEEEHRMFLLGLQKLGKGDWRGIARNYVISRTPTQVASHAQKYFIRQSN 658
Query: 183 TPKRRKRASIHDLTID---DTDLVPQQ----NQVP 210
+R++R+S+ D+ D DT +V Q N++P
Sbjct: 659 VSRRKRRSSLFDIVADEAADTAMVQQDFLSANELP 763
>TC228766 similar to GB|AAS99672.1|46518381|BT012528 At5g56840 {Arabidopsis
thaliana;} , partial (54%)
Length = 1220
Score = 89.4 bits (220), Expect = 3e-18
Identities = 39/70 (55%), Positives = 56/70 (79%)
Frame = +3
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ +K V WT++EHR FL GL+K GKG W+ ISR F+ ++TPTQ+ASHAQKY+LRL + K
Sbjct: 390 ERKKGVPWTEEEHRIFLVGLEKLGKGDWRGISRNFVTTRTPTQVASHAQKYFLRLATMDK 569
Query: 186 RRKRASIHDL 195
+++R+S+ DL
Sbjct: 570 KKRRSSLFDL 599
>TC227468 similar to UP|Q8H1D0 (Q8H1D0) Transcription factor MYBS3, partial
(56%)
Length = 1404
Score = 89.4 bits (220), Expect = 3e-18
Identities = 42/94 (44%), Positives = 62/94 (65%)
Frame = +3
Query: 128 EKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRR 187
+K V WT++EHR FL GL+K GKG W+ I+R F+ S+TPTQ+ASHAQKY++R + +R+
Sbjct: 384 KKGVPWTEEEHRLFLIGLQKLGKGDWRGIARNFVVSRTPTQVASHAQKYFIRQSHATRRK 563
Query: 188 KRASIHDLTIDDTDLVPQQNQVPQLNAPMQELSP 221
+R+S+ D+ D + P + L P Q P
Sbjct: 564 RRSSLFDMVPDMSSDQPSVPEEQVLLPPSQNSQP 665
>TC219288 similar to GB|AAS99672.1|46518381|BT012528 At5g56840 {Arabidopsis
thaliana;} , partial (41%)
Length = 720
Score = 87.8 bits (216), Expect = 9e-18
Identities = 37/68 (54%), Positives = 55/68 (80%)
Frame = +2
Query: 128 EKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPKRR 187
+K V WT++EHR FL GL+K GKG W+ ISR ++ S+TPTQ+ASHAQKY++RL + K++
Sbjct: 299 KKGVPWTEEEHRTFLVGLEKLGKGDWRGISRNYVTSRTPTQVASHAQKYFIRLATMNKKK 478
Query: 188 KRASIHDL 195
+R+S+ D+
Sbjct: 479 RRSSLFDM 502
>TC214991 similar to GB|AAS09980.1|41618916|AY519510 MYB transcription factor
{Arabidopsis thaliana;} , partial (33%)
Length = 677
Score = 87.4 bits (215), Expect = 1e-17
Identities = 36/73 (49%), Positives = 58/73 (79%)
Frame = +2
Query: 126 QNEKRVLWTDDEHRNFLRGLKKYGKGQWQTISREFLPSKTPTQIASHAQKYYLRLNSTPK 185
+ ++ V WT++EH+ FL GL+K GKG W+ IS+ ++ ++TPTQ+ASHAQKY+LR ++ +
Sbjct: 71 ERKRGVPWTEEEHKLFLVGLQKVGKGDWRGISKNYVKTRTPTQVASHAQKYFLRRSNLNR 250
Query: 186 RRKRASIHDLTID 198
RR+R+S+ D+T D
Sbjct: 251 RRRRSSLFDITTD 289
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,912,660
Number of Sequences: 63676
Number of extensions: 373091
Number of successful extensions: 2067
Number of sequences better than 10.0: 151
Number of HSP's better than 10.0 without gapping: 1564
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2015
length of query: 471
length of database: 12,639,632
effective HSP length: 101
effective length of query: 370
effective length of database: 6,208,356
effective search space: 2297091720
effective search space used: 2297091720
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0213.5


