

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
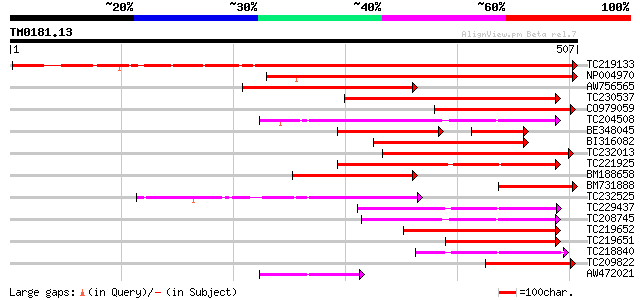
Query= TM0181.13
(507 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219133 UP|Q39879 (Q39879) Mitotic cyclin a2-type, complete 733 0.0
NP004970 mitotic cyclin a1-type 281 4e-76
AW756565 similar to PIR|T07675|T07 cyclin a2-type mitosis-speci... 238 6e-63
TC230537 mitotic cyclin a2-type 216 1e-56
CO979059 197 1e-50
TC204508 homologue to UP|Q39855 (Q39855) Cyclin, partial (98%) 186 2e-47
BE348045 similar to PIR|T07672|T07 cyclin a2-type mitosis-speci... 138 7e-42
BI316082 similar to PIR|T07672|T07 cyclin a2-type mitosis-speci... 166 2e-41
TC232013 similar to UP|Q9S957 (Q9S957) Cyclin A homolog, partial... 162 3e-40
TC221925 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type,... 150 1e-36
BM188658 similar to PIR|T07669|T07 cyclin a1-type mitosis-speci... 149 4e-36
BM731888 homologue to PIR|T07675|T076 cyclin a2-type mitosis-sp... 141 6e-34
TC232525 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type,... 133 2e-31
TC229437 similar to UP|Q40670 (Q40670) Cyclin (Fragment), partia... 129 3e-30
TC208745 homologue to UP|CG22_SOYBN (P25012) G2/mitotic-specific... 125 6e-29
TC219652 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, p... 114 1e-25
TC219651 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, p... 91 1e-18
TC218840 similar to UP|CG1B_MEDVA (P46277) G2/mitotic-specific c... 83 3e-16
TC209822 similar to UP|Q9M4K9 (Q9M4K9) Cyclin A3.1, partial (24%) 79 4e-15
AW472021 similar to PIR|S56679|S56 mitosis-specific cyclin CycII... 69 5e-12
>TC219133 UP|Q39879 (Q39879) Mitotic cyclin a2-type, complete
Length = 1804
Score = 733 bits (1892), Expect = 0.0
Identities = 392/509 (77%), Positives = 426/509 (83%), Gaps = 4/509 (0%)
Frame = +2
Query: 3 SNNHRRSSFSSSTASSLAKRHASDNNNNNNNNNNYVVGAAAVKAPAAASHSAKKRPPLSN 62
S +RRSSFSSST SSLAKRHAS + ++ AA ++ AKKRPPLSN
Sbjct: 47 STQNRRSSFSSSTTSSLAKRHASASTTSS--------------LAAAPTNMAKKRPPLSN 184
Query: 63 LTNHIASRNSSSQSLVPCVSKFAKTKKEAPPKVA----PALPNVKSAAAAVVFPKVATAT 118
LTN +A RNSSS VPC +KFAKTKK+ P + P L NVK A+A V+FPK
Sbjct: 185 LTNTVAHRNSSSS--VPCAAKFAKTKKDVVPACSGNKRPTLANVKPASA-VIFPKAN--- 346
Query: 119 ASFTERNEDDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDESMSSCDSFKSPAEIEY 178
SF +NE A V+VP +DVSP KSD SVS DESMSSCDSFKSP +IEY
Sbjct: 347 -SFPVKNEAPPTPPPPVATVTVPAPV-VDVSPSKSDTMSVSTDESMSSCDSFKSP-DIEY 517
Query: 179 VDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVNIDNDHMDPQL 238
VDNSDV+AVDSIERKTFS LNISD+ E GNICSR+ILVE LEKG+K VN+DN++ DPQL
Sbjct: 518 VDNSDVAAVDSIERKTFSHLNISDSTE--GNICSREILVE-LEKGDKFVNVDNNYADPQL 688
Query: 239 CASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLY 298
CA+FACDIYKHLRASEAKKRPSTDFMEK+QK+IN+SMRAILIDWLVEVAEEYRLVPDTLY
Sbjct: 689 CATFACDIYKHLRASEAKKRPSTDFMEKIQKEINSSMRAILIDWLVEVAEEYRLVPDTLY 868
Query: 299 LTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQ 358
LTVNYIDRYLSGN M+RQ+LQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQ
Sbjct: 869 LTVNYIDRYLSGNVMNRQRLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQ 1048
Query: 359 MESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCY 418
MES VLNFLKFEMTAPT+KCFLRRFVRAAQGV+EVPSLQLE LTNYIAELSLMEYSML Y
Sbjct: 1049MESAVLNFLKFEMTAPTVKCFLRRFVRAAQGVDEVPSLQLECLTNYIAELSLMEYSMLGY 1228
Query: 419 APSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPA 478
APSLVAASAIFLAKFILFPS KPW+STLQHYTLYQPSDLCVCVK+LHRL CNSPNSNLPA
Sbjct: 1229APSLVAASAIFLAKFILFPSKKPWNSTLQHYTLYQPSDLCVCVKDLHRLCCNSPNSNLPA 1408
Query: 479 IKEKYSQHKYKYVAKKYCPPSIPSEFFQN 507
I+EKYSQHKYKYVAKKYCPPSIP EFFQN
Sbjct: 1409IREKYSQHKYKYVAKKYCPPSIPPEFFQN 1495
>NP004970 mitotic cyclin a1-type
Length = 1047
Score = 281 bits (720), Expect = 4e-76
Identities = 139/280 (49%), Positives = 201/280 (71%), Gaps = 2/280 (0%)
Frame = +1
Query: 230 DNDHMDPQLCASFACDIYKHLRASEA--KKRPSTDFMEKVQKDINTSMRAILIDWLVEVA 287
+N PQL S+ DI+++LR E K+RP +++EK QK + +MR IL+DWLVEVA
Sbjct: 202 NNTLSSPQLDGSYVSDIHEYLREMEMQKKRRPMVNYIEKFQKIVTPTMRGILVDWLVEVA 381
Query: 288 EEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYIT 347
EEY+L+ DTL+L+V+YIDR+LS N +++ +LQLLGV+SM+IA+KYEE P V+EFC IT
Sbjct: 382 EEYKLLSDTLHLSVSYIDRFLSVNPVTKSRLQLLGVSSMLIAAKYEETDPPSVDEFCSIT 561
Query: 348 DNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAE 407
DNTY K EV++ME+++L LKFEM PT+ FLRR+ A V++ P+ Q+E L +YI E
Sbjct: 562 DNTYDKAEVVKMEADILKSLKFEMGNPTVSTFLRRYANVASDVQKTPNSQIEHLGSYIGE 741
Query: 408 LSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRL 467
LSL++Y L + PS+VAAS IFLAKFI++P + PW+S+L + Y+P++L CV LH L
Sbjct: 742 LSLLDYDCLRFLPSIVAASVIFLAKFIIWPEVHPWTSSLCECSGYKPAELKECVLILHDL 921
Query: 468 FCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEFFQN 507
+ + ++ A++EKY K+K VA PP +PS +F++
Sbjct: 922 YLSRKAASFKAVREKYKHQKFKCVANLPTPPYVPSCYFED 1041
>AW756565 similar to PIR|T07675|T07 cyclin a2-type mitosis-specific -
soybean, partial (28%)
Length = 472
Score = 238 bits (606), Expect = 6e-63
Identities = 116/156 (74%), Positives = 136/156 (86%)
Frame = +2
Query: 209 NICSRDILVEELEKGEKIVNIDNDHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQ 268
N+CSR+I+VE E+ +KIVNIDN + D QLCA++ CDIYKHLR S KKRPSTDFM+ +Q
Sbjct: 2 NVCSREIIVELEERVDKIVNIDNIYSDTQLCATYVCDIYKHLRES*EKKRPSTDFMDTIQ 181
Query: 269 KDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMI 328
KDIN SMR IL+DWLVEVAEEYRLVP+TLYLTVNY+DR LS NAM+RQ++QLLGV+ MMI
Sbjct: 182 KDINVSMRTILVDWLVEVAEEYRLVPETLYLTVNYLDRSLSWNAMNRQRIQLLGVSCMMI 361
Query: 329 ASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVL 364
ASKYEEICAP VEE+ YITD+TY KEEVLQMES V+
Sbjct: 362 ASKYEEICAPHVEEYRYITDDTYLKEEVLQMESCVV 469
>TC230537 mitotic cyclin a2-type
Length = 993
Score = 216 bits (551), Expect = 1e-56
Identities = 110/193 (56%), Positives = 143/193 (73%)
Frame = +1
Query: 300 TVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQM 359
+VN IDRYLS + +QKLQLLGV M+IASKYEE+CAP+VEEFC+ITDNTY KEEVL+M
Sbjct: 7 SVNLIDRYLSTRLIQKQKLQLLGVTCMLIASKYEEMCAPRVEEFCFITDNTYTKEEVLKM 186
Query: 360 ESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYA 419
E EVL+ + F+++ PTIK FLRRF++AAQ + P ++LE L NY+AEL+L+E + +
Sbjct: 187 EREVLDLVHFQLSVPTIKTFLRRFIQAAQSSYKAPCVELEFLANYLAELALVECNFFQFL 366
Query: 420 PSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAI 479
PSLVAASA+FLAK+ L S PW+ TL+HYT Y+ S+L V L L N+ S+L A+
Sbjct: 367 PSLVAASAVFLAKWTLNESEHPWNPTLKHYTKYKASELKTVVLALQDLQLNTKGSSLNAV 546
Query: 480 KEKYSQHKYKYVA 492
EKY Q K+ VA
Sbjct: 547 PEKYKQQKFNCVA 585
>CO979059
Length = 766
Score = 197 bits (500), Expect = 1e-50
Identities = 97/127 (76%), Positives = 111/127 (87%), Gaps = 1/127 (0%)
Frame = -2
Query: 381 RRFVRAA-QGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSI 439
RRFVRAA V+E+ SLQLE LTN+IAELSL+EYSML Y PSL+AAS IFLA+FILFPS
Sbjct: 765 RRFVRAAAHDVQEIXSLQLEYLTNFIAELSLLEYSMLSYPPSLIAASVIFLARFILFPSK 586
Query: 440 KPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPS 499
KPW+STLQHYTLY+PSDLC CVK+LHRL C+S +SNLPAI++KYSQHKYK VAKK+ PPS
Sbjct: 585 KPWNSTLQHYTLYRPSDLCACVKDLHRLCCSSHDSNLPAIRDKYSQHKYKCVAKKHIPPS 406
Query: 500 IPSEFFQ 506
IP E FQ
Sbjct: 405 IPREVFQ 385
>TC204508 homologue to UP|Q39855 (Q39855) Cyclin, partial (98%)
Length = 1283
Score = 186 bits (472), Expect = 2e-47
Identities = 107/275 (38%), Positives = 166/275 (59%), Gaps = 6/275 (2%)
Frame = +2
Query: 224 EKIVNIDNDHMDPQLCA------SFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRA 277
E+I++ID +D +L A + DIYK + E + RP D++ Q +IN MRA
Sbjct: 251 EQIIDIDASDVDNELAAVELAAVEYIDDIYKFYKLVENESRPH-DYIGS-QPEINERMRA 424
Query: 278 ILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICA 337
IL+DWL++V ++ L +TLYLT+N IDR+L+ + R++LQL+G+++M++ASKYEEI
Sbjct: 425 ILVDWLIDVHTKFELSLETLYLTINIIDRFLAVKTVPRRELQLVGISAMLMASKYEEIWP 604
Query: 338 PQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQ 397
P+V +F ++D Y E++L ME +LN L++ +T PT FL RF++AA VP +
Sbjct: 605 PEVNDFVCLSDRAYTHEQILAMEKTILNKLEWTLTVPTPFVFLVRFIKAA-----VPDQE 769
Query: 398 LESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDL 457
LE++ ++++EL +M Y+ L Y PS+VAASA+F A+ L W+ TL+ +T Y L
Sbjct: 770 LENMAHFMSELGMMNYATLMYCPSMVAASAVFAARCTL-NKAPLWNETLKLHTGYSQEQL 946
Query: 458 CVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
C + L N L + KYS + VA
Sbjct: 947 MDCARLLVGFHSTLGNGKLRVVYRKYSDPQKGAVA 1051
>BE348045 similar to PIR|T07672|T07 cyclin a2-type mitosis-specific -
soybean, partial (34%)
Length = 491
Score = 138 bits (348), Expect(2) = 7e-42
Identities = 67/95 (70%), Positives = 80/95 (83%)
Frame = -1
Query: 294 PDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFK 353
PDTLYLTVN IDR LS + + +Q+LQLLGV M+IASKYEEICAP+VEEFC+ITDNTY K
Sbjct: 491 PDTLYLTVNLIDRSLSQSLVQKQRLQLLGVTCMLIASKYEEICAPRVEEFCFITDNTYTK 312
Query: 354 EEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQ 388
EVL+MESEVLN L F+++ PT K FLRRF+ A+Q
Sbjct: 311 AEVLKMESEVLNLLHFQLSVPTTKTFLRRFILASQ 207
Score = 50.8 bits (120), Expect(2) = 7e-42
Identities = 25/51 (49%), Positives = 35/51 (68%)
Frame = -2
Query: 414 SMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKEL 464
S L + PSL+AASA+ LA++ L S PW+ST++HYT Y+ S+L V L
Sbjct: 196 SFLQFLPSLIAASAVLLARWTLNQSEHPWNSTMEHYTNYKVSELKTTVLAL 44
>BI316082 similar to PIR|T07672|T07 cyclin a2-type mitosis-specific -
soybean, partial (31%)
Length = 456
Score = 166 bits (421), Expect = 2e-41
Identities = 82/139 (58%), Positives = 108/139 (76%)
Frame = +3
Query: 326 MMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVR 385
M+IASKYEEICAP+VEEFC+ITDNTY K EVL+MESEVLN L F+++ PT K FLRRF+
Sbjct: 6 MLIASKYEEICAPRVEEFCFITDNTYTKAEVLKMESEVLNLLHFQLSVPTTKTFLRRFIL 185
Query: 386 AAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSST 445
A++ +V ++LE L NY+AEL+L+EYS L + PSL+AASA+ LA++ L S PW+ST
Sbjct: 186 ASESSYKVSYVELEFLANYLAELTLVEYSFLQFLPSLIAASAVLLARWTLNQSEHPWNST 365
Query: 446 LQHYTLYQPSDLCVCVKEL 464
++HYT Y+ S+L V L
Sbjct: 366 MEHYTNYKVSELKTTVLAL 422
>TC232013 similar to UP|Q9S957 (Q9S957) Cyclin A homolog, partial (38%)
Length = 814
Score = 162 bits (410), Expect = 3e-40
Identities = 84/171 (49%), Positives = 116/171 (67%)
Frame = +1
Query: 334 EICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEV 393
EI AP++E+FC+ITDNTY K EVL+ME +VL ++++ APTI+ F+RRF+RAAQ +
Sbjct: 1 EINAPRIEDFCFITDNTYTKAEVLKMERQVLKSSEYQLFAPTIQTFVRRFLRAAQASYKD 180
Query: 394 PSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQ 453
SL+LE L NY+AEL+LM+Y L + PS++AASA+FLA++ L S PW+ TLQHY Y+
Sbjct: 181 QSLELEYLANYLAELTLMDYGFLNFLPSIIAASAVFLARWTLDQSNHPWNPTLQHYACYK 360
Query: 454 PSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCPPSIPSEF 504
SDL V L L N+ L A++ KY Q K+K VA P + + F
Sbjct: 361 ASDLKTTVLALQDLQLNTDGCPLTAVRTKYRQDKFKCVAALSSPKLLETLF 513
>TC221925 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type, partial
(46%)
Length = 802
Score = 150 bits (380), Expect = 1e-36
Identities = 84/200 (42%), Positives = 129/200 (64%), Gaps = 1/200 (0%)
Frame = +2
Query: 294 PDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFK 353
P+TLYLT+ +DR+LS A+ R++LQL+G++SM+IASKYEEI AP+V +F I+DN Y
Sbjct: 2 PETLYLTLXIVDRFLSVKAVPRRELQLVGISSMLIASKYEEIWAPEVNDFECISDNAYVS 181
Query: 354 EEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEY 413
++VL ME +L L++ +T PT FL R+++A+ ++ ++E++ ++AEL LM Y
Sbjct: 182 QQVLMMEXXILRKLEWYLTVPTPYHFLVRYIKASTPSDK----EMENMVFFLAELGLMHY 349
Query: 414 -SMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSP 472
+ + Y PSL+AA+A+F A+ L S W+STL+HYT Y L C K + L +P
Sbjct: 350 PTAILYRPSLIAAAAVFAARCTLGRS-PFWTSTLKHYTGYSEEQLRDCAKIMVNLHAAAP 526
Query: 473 NSNLPAIKEKYSQHKYKYVA 492
S L A+ +K+ VA
Sbjct: 527 GSKLRAVYKKFCNSDLSAVA 586
>BM188658 similar to PIR|T07669|T07 cyclin a1-type mitosis-specific -
soybean, partial (32%)
Length = 438
Score = 149 bits (375), Expect = 4e-36
Identities = 68/111 (61%), Positives = 95/111 (85%)
Frame = +3
Query: 254 EAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVAEEYRLVPDTLYLTVNYIDRYLSGNAM 313
+ K+RP D+M+KVQK + T+MR IL+DWLVEVAEEY+L+ DTL+L+V+YIDR+LS N +
Sbjct: 102 QRKRRPMIDYMDKVQKQVTTTMRTILVDWLVEVAEEYKLLSDTLHLSVSYIDRFLSVNPV 281
Query: 314 SRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVL 364
S+ +LQLLGV+SM+IA+KYEE+ P+V+ FC ITDNTY K EV++ME+++L
Sbjct: 282 SKSRLQLLGVSSMLIAAKYEEVDPPRVDPFCNITDNTYHKAEVVKMEADML 434
>BM731888 homologue to PIR|T07675|T076 cyclin a2-type mitosis-specific -
soybean, partial (14%)
Length = 422
Score = 141 bits (356), Expect = 6e-34
Identities = 62/70 (88%), Positives = 67/70 (95%)
Frame = +2
Query: 438 SIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVAKKYCP 497
S KPW+STLQHYTLY+PSDLCVCV++LHRL CNSPNSNLPAI+EKYSQHKYKYVAKKYCP
Sbjct: 83 SKKPWTSTLQHYTLYKPSDLCVCVRDLHRLCCNSPNSNLPAIREKYSQHKYKYVAKKYCP 262
Query: 498 PSIPSEFFQN 507
PSIP EFFQN
Sbjct: 263 PSIPPEFFQN 292
>TC232525 homologue to UP|Q39880 (Q39880) Mitotic cyclin b1-type, partial
(73%)
Length = 1242
Score = 133 bits (334), Expect = 2e-31
Identities = 93/262 (35%), Positives = 149/262 (56%), Gaps = 6/262 (2%)
Frame = +1
Query: 114 VATATASFTERNEDDAAAAASGARVSVPVSTSMDVSPCKSDGCSVSMDES-----MSSCD 168
+A A A+ TE+N+ + +GA V+ + P + G + E +SS D
Sbjct: 457 LANAQAA-TEKNKKSSTEVNNGAVVATDGVGVGNFVPARKVGAAKKPKEEPEVIVISSDD 633
Query: 169 SFKSPAEIEYVDNSDVSAVDSIERKTFSILNISDAKEQAGNICSRDILVEELEKGEKIVN 228
+ + + SA + K FS ++ A+ +A RD+ ++N
Sbjct: 634 ESEEKQAAKGKKEREKSARKNA--KAFS--SVLSARSKAACGLPRDL----------VMN 771
Query: 229 IDNDHMDPQLCAS-FACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDWLVEVA 287
ID MD +L A+ + DIYK + +E ++ D+M Q DIN MR+IL+DWL+EV
Sbjct: 772 IDATDMDNELAAAEYIDDIYKFYKETE-EEGCVHDYMGS-QPDINAKMRSILVDWLIEVH 945
Query: 288 EEYRLVPDTLYLTVNYIDRYLSGNAMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYIT 347
++ L+P+TLYLT+N +DR+LS A+ R++LQL+G++SM+IASKYEEI AP+V +F I+
Sbjct: 946 RKFELMPETLYLTLNIVDRFLSVKAVPRRELQLVGISSMLIASKYEEIWAPEVNDFECIS 1125
Query: 348 DNTYFKEEVLQMESEVLNFLKF 369
DN Y ++VL ME ++L L++
Sbjct: 1126DNAYVSQQVLMMEKKILRKLEW 1191
>TC229437 similar to UP|Q40670 (Q40670) Cyclin (Fragment), partial (77%)
Length = 830
Score = 129 bits (324), Expect = 3e-30
Identities = 72/182 (39%), Positives = 109/182 (59%)
Frame = +1
Query: 312 AMSRQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEM 371
A+ R+KLQL+GV +M+IA KYEE+ P VE+F ITD Y + EVL ME ++N L+F++
Sbjct: 10 AVIRKKLQLVGVTAMLIACKYEEVSVPTVEDFILITDKAYTRNEVLDMEKLMMNILQFKL 189
Query: 372 TAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLA 431
+ PT F+RRF++AA +LE L+ ++ EL L+E ML ++PSL+AA+AI+ A
Sbjct: 190 SVPTPYMFMRRFLKAAHS-----DKKLELLSFFLVELCLVECKMLKFSPSLLAAAAIYTA 354
Query: 432 KFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYV 491
+ L+ K W+ T + YT Y L C + + + + L + KY+ KY
Sbjct: 355 QCSLY-QFKQWTKTTEWYTDYSEEKLLECSRLMVTFHQKAGSGKLTGVYRKYNTWKYGCA 531
Query: 492 AK 493
AK
Sbjct: 532 AK 537
>TC208745 homologue to UP|CG22_SOYBN (P25012) G2/mitotic-specific cyclin
S13-7 (B-like cyclin) (Fragment), partial (74%)
Length = 879
Score = 125 bits (313), Expect = 6e-29
Identities = 69/178 (38%), Positives = 107/178 (59%)
Frame = +1
Query: 315 RQKLQLLGVASMMIASKYEEICAPQVEEFCYITDNTYFKEEVLQMESEVLNFLKFEMTAP 374
R++LQL+G+++M++ASKYEEI P+V +F ++D Y E++L ME +LN L++ +T P
Sbjct: 7 RRELQLVGISAMLMASKYEEIWPPEVNDFVCLSDRAYTHEQILAMEKTILNKLEWTLTVP 186
Query: 375 TIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFI 434
T FL RF++AA VP +LE++ ++++EL +M Y+ L Y PS+VAASA+F A+
Sbjct: 187 TPFVFLVRFIKAA-----VPDQELENMAHFMSELGMMNYATLMYCPSMVAASAVFAARCT 351
Query: 435 LFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
L W+ TL+ +T Y L C + L N L + KYS + VA
Sbjct: 352 L-NKAPLWNETLKLHTGYSQEQLMDCARLLVGFHSTLENGKLRVVYRKYSDPQKGAVA 522
>TC219652 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, partial (30%)
Length = 670
Score = 114 bits (284), Expect = 1e-25
Identities = 60/140 (42%), Positives = 91/140 (64%)
Frame = +1
Query: 353 KEEVLQMESEVLNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLME 412
+E + E E F+++ PT K FLRRF+ A+Q +V ++LE L NY+AEL+L+E
Sbjct: 1 RERERERERERERERDFQLSVPTTKTFLRRFILASQSSYKVSYVELEFLANYLAELTLVE 180
Query: 413 YSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSP 472
YS L + PSL+AASA+ LA++ L S PW+ST++HYT Y+ S+L V L L +
Sbjct: 181 YSFLQFLPSLIAASAVLLARWTLNQSEHPWNSTMEHYTNYKVSELKTAVLALADLLHDMK 360
Query: 473 NSNLPAIKEKYSQHKYKYVA 492
++ +I+EKY+Q K++ VA
Sbjct: 361 GCSVHSIREKYTQQKFRSVA 420
>TC219651 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, partial (24%)
Length = 701
Score = 90.9 bits (224), Expect = 1e-18
Identities = 48/103 (46%), Positives = 70/103 (67%)
Frame = +2
Query: 390 VEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLVAASAIFLAKFILFPSIKPWSSTLQHY 449
V +V ++LE L NY+AEL+L+EYS L + PSL+AASA+ LA++ L S PW+ST++HY
Sbjct: 107 VIQVSYVELEFLANYLAELTLVEYSFLQFLPSLIAASAVLLARWTLNQSEHPWNSTMEHY 286
Query: 450 TLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQHKYKYVA 492
T Y+ S+L V L L + +L +I+EKY Q K++ VA
Sbjct: 287 TNYKVSELKTTVLALADLQHDMKGCSLNSIREKYKQQKFRSVA 415
>TC218840 similar to UP|CG1B_MEDVA (P46277) G2/mitotic-specific cyclin 1
(B-like cyclin) (CycMs1), partial (32%)
Length = 922
Score = 83.2 bits (204), Expect = 3e-16
Identities = 46/136 (33%), Positives = 74/136 (53%)
Frame = +1
Query: 364 LNFLKFEMTAPTIKCFLRRFVRAAQGVEEVPSLQLESLTNYIAELSLMEYSMLCYAPSLV 423
+N L+F M+ PT F++RF++AAQ +LE L ++ EL+L+EY ML + PSL+
Sbjct: 1 VNLLQFNMSVPTAYVFMKRFLKAAQA-----DRKLELLAFFLVELTLVEYEMLKFPPSLL 165
Query: 424 AASAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKY 483
AA+A++ A+ ++ K WS T + ++ Y L C + + N L + KY
Sbjct: 166 AAAAVYTAQCTIY-GFKQWSKTCEWHSNYSEDQLLECSTLMAAFHQKAGNGKLTGVHRKY 342
Query: 484 SQHKYKYVAKKYCPPS 499
K+ Y AK C P+
Sbjct: 343 CSSKFSYTAK--CEPA 384
>TC209822 similar to UP|Q9M4K9 (Q9M4K9) Cyclin A3.1, partial (24%)
Length = 640
Score = 79.3 bits (194), Expect = 4e-15
Identities = 36/81 (44%), Positives = 50/81 (61%)
Frame = +2
Query: 426 SAIFLAKFILFPSIKPWSSTLQHYTLYQPSDLCVCVKELHRLFCNSPNSNLPAIKEKYSQ 485
S +FLA+F+ PW+S L T Y+P+DL CV LH L+ + ++L A++EKY Q
Sbjct: 2 SVVFLARFMFSTKTHPWNSALHQLTRYKPADLKECVLNLHDLYLSRRGASLQAVREKYKQ 181
Query: 486 HKYKYVAKKYCPPSIPSEFFQ 506
HK+K VA PP IP FF+
Sbjct: 182 HKFKCVATTPSPPEIPLSFFE 244
>AW472021 similar to PIR|S56679|S56 mitosis-specific cyclin CycIII - alfalfa,
partial (24%)
Length = 317
Score = 68.9 bits (167), Expect = 5e-12
Identities = 40/95 (42%), Positives = 57/95 (59%), Gaps = 1/95 (1%)
Frame = +2
Query: 224 EKIVNIDN-DHMDPQLCASFACDIYKHLRASEAKKRPSTDFMEKVQKDINTSMRAILIDW 282
E +++ID D +P + D+Y H R E S D+M + Q DIN MRAILIDW
Sbjct: 35 ETVLDIDTCDANNPLAVVDYIEDLYAHYRKMEGTSCVSPDYMAQ-QFDINERMRAILIDW 211
Query: 283 LVEVAEEYRLVPDTLYLTVNYIDRYLSGNAMSRQK 317
L+E ++ L+ +TL LTVN IDR+L+ + R+K
Sbjct: 212 LIEGHDKLDLLHETLVLTVNLIDRFLAKQTVVRKK 316
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.315 0.127 0.359
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,950,559
Number of Sequences: 63676
Number of extensions: 309668
Number of successful extensions: 4985
Number of sequences better than 10.0: 211
Number of HSP's better than 10.0 without gapping: 2801
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3695
length of query: 507
length of database: 12,639,632
effective HSP length: 101
effective length of query: 406
effective length of database: 6,208,356
effective search space: 2520592536
effective search space used: 2520592536
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0181.13


