

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0132.1
(267 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
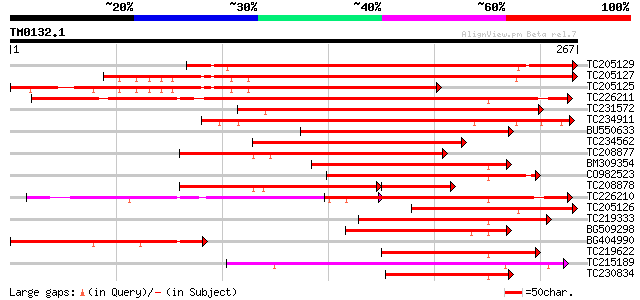
Sequences producing significant alignments: (bits) Value
TC205129 homologue to UP|HT22_ARATH (P46604) Homeobox-leucine zi... 325 1e-89
TC205127 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipp... 323 5e-89
TC205125 similar to UP|HAT9_ARATH (P46603) Homeobox-leucine zipp... 238 2e-63
TC226211 UP|Q39862 (Q39862) Homeobox-leucine zipper protein, com... 206 7e-54
TC231572 similar to UP|HT14_ARATH (P46665) Homeobox-leucine zipp... 205 2e-53
TC234911 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipp... 197 4e-51
BU550633 similar to PIR|T52367|T52 homeobox protein HAT14 [impor... 166 1e-41
TC234562 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipp... 159 2e-39
TC208877 similar to UP|Q39862 (Q39862) Homeobox-leucine zipper p... 149 1e-36
BM309354 similar to PIR|T06438|T06 homeobox-leucine zipper prote... 147 4e-36
CO982523 143 7e-35
TC208878 similar to UP|Q39862 (Q39862) Homeobox-leucine zipper p... 96 8e-34
TC226210 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper... 129 1e-30
TC205126 similar to UP|HAT9_ARATH (P46603) Homeobox-leucine zipp... 128 2e-30
TC219333 homologue to UP|Q42437 (Q42437) HD-ZIP protein, partial... 124 6e-29
BG509298 weakly similar to PIR|T09784|T097 homeobox leucine zipp... 107 7e-24
BG404990 105 3e-23
TC219622 similar to UP|Q6I664 (Q6I664) HD-ZIP protein (Fragment)... 97 8e-21
TC215189 similar to UP|Q39941 (Q39941) HAHB-1, partial (70%) 94 5e-20
TC230834 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper... 93 1e-19
>TC205129 homologue to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (47%)
Length = 793
Score = 325 bits (834), Expect = 1e-89
Identities = 168/191 (87%), Positives = 176/191 (91%), Gaps = 7/191 (3%)
Frame = +3
Query: 84 SFSTARVVKRERVDLSCE----EIEAEERLSSRVSDEDDDGTNARKKLRLTKEQSALLEE 139
SFST RV+KRER DLSCE E+EAEER+SSRVSDED+DGTNARKKLRLTKEQSALLEE
Sbjct: 3 SFSTGRVIKRER-DLSCEDIEVEVEAEERVSSRVSDEDEDGTNARKKLRLTKEQSALLEE 179
Query: 140 SFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDE 199
SFKQHSTLNPKQKQALARQLNLR RQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDE
Sbjct: 180 SFKQHSTLNPKQKQALARQLNLRPRQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDE 359
Query: 200 NRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERLG---DGGSNIKSPFTITPKPHF 256
NRRLKKELQELKALKLAQPLYMPM AATLTMCPSCERLG D GSN KSPF++ PKPHF
Sbjct: 360 NRRLKKELQELKALKLAQPLYMPMPAATLTMCPSCERLGGVSDNGSN-KSPFSMAPKPHF 536
Query: 257 FNPFTHPSAAC 267
+NPF +PSAAC
Sbjct: 537 YNPFANPSAAC 569
>TC205127 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (59%)
Length = 1036
Score = 323 bits (828), Expect = 5e-89
Identities = 184/257 (71%), Positives = 201/257 (77%), Gaps = 34/257 (13%)
Frame = +3
Query: 45 PYSSKE--PSLTLGLS---------------GNNKVY-CEDPLE-LSRQT-SPHSDVVSS 84
PY+S E PSLTLGLS NNKV C+DPL+ LS QT SPH VSS
Sbjct: 21 PYNSSEAEPSLTLGLSRESYLKVPKNIIGQNSNNKVSSCDDPLDHLSTQTNSPHHSAVSS 200
Query: 85 FSTARVVKRERVDLSCEEI----EAEERLSS-----RVSDEDDDGTNARKKLRLTKEQSA 135
FS+ RV KRER DLSCEE+ E ++R S R +DED+DGT ARKKLRL+KEQSA
Sbjct: 201 FSSGRV-KRER-DLSCEEVVDATEIDQRDHSCEGIVRATDEDEDGTAARKKLRLSKEQSA 374
Query: 136 LLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCET 195
LLEESFKQHSTLNPKQKQALA+QLNLR RQVEVWFQNRRARTKLKQTEVDCEFLKKCCET
Sbjct: 375 LLEESFKQHSTLNPKQKQALAKQLNLRPRQVEVWFQNRRARTKLKQTEVDCEFLKKCCET 554
Query: 196 LTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERL-----GDGGSNIKSPFTI 250
LTDENRRL+KELQELKALKLAQPLYMPM AATLTMCPSCERL G GG + K+PF++
Sbjct: 555 LTDENRRLQKELQELKALKLAQPLYMPMPAATLTMCPSCERLGGGINGGGGGSPKTPFSM 734
Query: 251 TPKPHFFNPFTHPSAAC 267
PKPHFFNPF +PSAAC
Sbjct: 735 APKPHFFNPFANPSAAC 785
>TC205125 similar to UP|HAT9_ARATH (P46603) Homeobox-leucine zipper protein
HAT9 (Homeodomain-leucine zipper protein HAT9)
(Homeodomain transcription factor HAT9) (HD-ZIP protein
9), partial (38%)
Length = 813
Score = 238 bits (608), Expect = 2e-63
Identities = 155/251 (61%), Positives = 168/251 (66%), Gaps = 48/251 (19%)
Frame = +3
Query: 1 MGLDHDAS-NPGLHLALGLSLTTTNTSKETTTTTTTSSP------------------KPT 41
MGLD DAS N GLHL LGL+LT T TTTT S P +PT
Sbjct: 87 MGLDQDASSNSGLHLILGLALTAT-------TTTTPSLPSISNKLDHVDHHHHLITLRPT 245
Query: 42 VMKPYSSKE--PSLTLGLS---------------GNNKVY-CEDPLE-LSRQT-SPHSDV 81
PY+S E PSLTLGLS NNKV C+DPL+ LS QT SPH
Sbjct: 246 TKSPYNSSEAEPSLTLGLSRESYLKVPKNIIGQNSNNKVSSCDDPLDHLSTQTNSPHHSA 425
Query: 82 VSSFSTARVVKRERVDLSCEEI----EAEERLSS-----RVSDEDDDGTNARKKLRLTKE 132
VSSFS+ RV KRER DLSCEE+ E ++R S R +DED+DGT ARKKLRL+KE
Sbjct: 426 VSSFSSGRV-KRER-DLSCEEVVDATEIDQRDHSCEGIVRATDEDEDGTAARKKLRLSKE 599
Query: 133 QSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKC 192
QSALLEESFKQHSTLNP QALA+QLNLR RQVEVWFQNRRARTKLKQTEVDCEFLKKC
Sbjct: 600 QSALLEESFKQHSTLNPNFIQALAKQLNLRPRQVEVWFQNRRARTKLKQTEVDCEFLKKC 779
Query: 193 CETLTDENRRL 203
CETLTDENRRL
Sbjct: 780 CETLTDENRRL 812
>TC226211 UP|Q39862 (Q39862) Homeobox-leucine zipper protein, complete
Length = 1249
Score = 206 bits (525), Expect = 7e-54
Identities = 125/256 (48%), Positives = 161/256 (62%), Gaps = 1/256 (0%)
Frame = +3
Query: 11 GLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTLGLSGNNKVYCEDPLE 70
GL L+L L +++ S + + P T S + S G+ N D E
Sbjct: 162 GLSLSLSFPLLSSSPSSHNPQKPSWNDPIFTS----SGEAGSFLRGIDVNRLPSVVDCEE 329
Query: 71 LSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKLRLT 130
+ +SP+S VSS S KR + + EE + + S + +++D +RKKLRL+
Sbjct: 330 EAGVSSPNS-TVSSVSG----KRSERETNGEENDTDRACSRGIISDEEDAETSRKKLRLS 494
Query: 131 KEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLK 190
K+QS +LEESFK+H+TLNPKQK ALA+QL LRARQVEVWFQNRRARTKLKQTEVDCEFLK
Sbjct: 495 KDQSIVLEESFKEHNTLNPKQKLALAKQLGLRARQVEVWFQNRRARTKLKQTEVDCEFLK 674
Query: 191 KCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNIKSPFT 249
+CCE LT+ENRRL+KE+QEL+ALKL+ YM M+ TLTMCPSCER+ S+ P T
Sbjct: 675 RCCENLTEENRRLQKEVQELRALKLSPQFYMHMTPPTTLTMCPSCERVAVPPSSAVDPAT 854
Query: 250 ITPKPHFFNPFTHPSA 265
H P +HP A
Sbjct: 855 ----RHHHVPPSHPRA 890
>TC231572 similar to UP|HT14_ARATH (P46665) Homeobox-leucine zipper protein
HAT14 (HD-ZIP protein 14), partial (53%)
Length = 835
Score = 205 bits (521), Expect = 2e-53
Identities = 102/148 (68%), Positives = 119/148 (79%), Gaps = 4/148 (2%)
Frame = +3
Query: 108 RLSSRVSDEDDD----GTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRA 163
R SSR SD+DD+ G N RKKLRL+KEQSA LEESFK+H+TLNPKQK ALA+QLNL+
Sbjct: 3 RTSSRASDDDDNNNGSGGNTRKKLRLSKEQSAFLEESFKEHNTLNPKQKLALAKQLNLQP 182
Query: 164 RQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPM 223
RQVEVWF NRRARTKLKQTEVDCE+LK+CCETLT+ENRRL KELQEL+ALK + P YM +
Sbjct: 183 RQVEVWFFNRRARTKLKQTEVDCEYLKRCCETLTEENRRLHKELQELRALKTSNPFYMQL 362
Query: 224 SAATLTMCPSCERLGDGGSNIKSPFTIT 251
A TLTMCPSCER+ ++ + T
Sbjct: 363 PATTLTMCPSCERVATNSTSTSLSISAT 446
>TC234911 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (40%)
Length = 799
Score = 197 bits (501), Expect = 4e-51
Identities = 113/196 (57%), Positives = 139/196 (70%), Gaps = 20/196 (10%)
Frame = +1
Query: 91 VKRERVD-LSCEEIEAE-ERLSSRVSDEDDDGTNARKKLRLTKEQSALLEESFKQHSTLN 148
+KRER L EE E E++ +RV + D+ N RKKLRLTKEQ+A+LEE+F++HSTLN
Sbjct: 7 IKRERDQVLGGEEFEVVVEKVPTRVVGDVDEDGNPRKKLRLTKEQAAVLEENFREHSTLN 186
Query: 149 PKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQ 208
PKQKQ LA +LNLRARQVEVWFQNRRARTKLKQTE DCE LKKCC+TLT EN++L+KELQ
Sbjct: 187 PKQKQELAMKLNLRARQVEVWFQNRRARTKLKQTESDCELLKKCCDTLTVENKKLQKELQ 366
Query: 209 ELKALKLAQ-PLYMPMSAATLTMCPSCERL-------GDGGSNIKSPFT---ITPKPHFF 257
ELK+++ PLYM + AATL++CPSCER+ GD +N S T I K H
Sbjct: 367 ELKSMQATPVPLYMQIPAATLSICPSCERICGGNENNGDNNNNASSHTTTLLIGSKTHHH 546
Query: 258 N-------PFTHPSAA 266
+ PF H S+A
Sbjct: 547 SFYKSNNYPFPHSSSA 594
>BU550633 similar to PIR|T52367|T52 homeobox protein HAT14 [imported] -
Arabidopsis thaliana (fragment), partial (61%)
Length = 694
Score = 166 bits (419), Expect = 1e-41
Identities = 78/100 (78%), Positives = 89/100 (89%)
Frame = -2
Query: 138 EESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLT 197
EESFK+H+TLN QK A A+QLNL RQVEVWFQNRRARTKLKQTEVDCE+LK+CCETLT
Sbjct: 693 EESFKEHTTLNXXQKLAXAKQLNLXXRQVEVWFQNRRARTKLKQTEVDCEYLKRCCETLT 514
Query: 198 DENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERL 237
+ENRRL+KELQEL+ALK +QP YM + A TLTMCPSCER+
Sbjct: 513 EENRRLQKELQELRALKSSQPFYMQLPATTLTMCPSCERV 394
>TC234562 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (33%)
Length = 585
Score = 159 bits (401), Expect = 2e-39
Identities = 78/101 (77%), Positives = 87/101 (85%)
Frame = +3
Query: 115 DEDDDGTNARKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRR 174
D ++ +RKKL+LTKEQSA LE+ FK HS+LNP QKQALA QLNL+ RQVEVWFQNRR
Sbjct: 270 DSNNSNNGSRKKLKLTKEQSATLEDIFKLHSSLNPAQKQALAEQLNLKHRQVEVWFQNRR 449
Query: 175 ARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKL 215
ARTKLKQTEVDCEFLKKCCE LTDEN RLKKELQEL+A K+
Sbjct: 450 ARTKLKQTEVDCEFLKKCCEKLTDENLRLKKELQELRAQKI 572
>TC208877 similar to UP|Q39862 (Q39862) Homeobox-leucine zipper protein,
partial (46%)
Length = 768
Score = 149 bits (377), Expect = 1e-36
Identities = 82/137 (59%), Positives = 99/137 (71%), Gaps = 11/137 (8%)
Frame = +3
Query: 81 VVSSFSTARVVKRERVDLSCEEIEAEERLSSRV---SDEDDDGT--------NARKKLRL 129
V S ST + +R + E E SSR SD+D+ G +RKKLRL
Sbjct: 357 VSSPNSTISSISGKRSEREGNGEENERTSSSRGGGGSDDDEGGACGGDADADASRKKLRL 536
Query: 130 TKEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFL 189
+KEQ+ +LEE+FK+H+TLNPKQKQALA+QLNL RQVEVWFQNRRARTKLKQTEVDCE+L
Sbjct: 537 SKEQALVLEETFKEHNTLNPKQKQALAKQLNLMPRQVEVWFQNRRARTKLKQTEVDCEYL 716
Query: 190 KKCCETLTDENRRLKKE 206
K+CCE LT+ENRRL+KE
Sbjct: 717 KRCCENLTEENRRLQKE 767
>BM309354 similar to PIR|T06438|T06 homeobox-leucine zipper protein homolog -
soybean (fragment), partial (58%)
Length = 428
Score = 147 bits (372), Expect = 4e-36
Identities = 71/95 (74%), Positives = 84/95 (87%), Gaps = 1/95 (1%)
Frame = +1
Query: 143 QHSTLNPKQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRR 202
+HSTLNPK+KQALA +LNL+ RQVEVWFQNRRARTKLKQTEVDCE+LK+C E LT+ENRR
Sbjct: 1 EHSTLNPKRKQALAEELNLKPRQVEVWFQNRRARTKLKQTEVDCEYLKRCYENLTEENRR 180
Query: 203 LKKELQELKALKLAQPLYMPMS-AATLTMCPSCER 236
L KE+QEL+ALKL+ +YM M+ TLT+CPSCER
Sbjct: 181 LHKEVQELRALKLSPQMYMHMNPPTTLTICPSCER 285
>CO982523
Length = 877
Score = 143 bits (361), Expect = 7e-35
Identities = 73/102 (71%), Positives = 84/102 (81%), Gaps = 1/102 (0%)
Frame = -2
Query: 150 KQKQALARQLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQE 209
K+KQALA +LNL+ RQVEVWFQNRRARTKLKQTEVDCE+LKKCCE LT+ENRRL KE+QE
Sbjct: 876 KRKQALAEELNLKPRQVEVWFQNRRARTKLKQTEVDCEYLKKCCENLTEENRRLHKEVQE 697
Query: 210 LKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNIKSPFTI 250
L+ALKL+ +YM M+ TLTMCPSCER S+ SP TI
Sbjct: 696 LRALKLSPQMYMHMNPPTTLTMCPSCERTHSSASS--SPATI 577
>TC208878 similar to UP|Q39862 (Q39862) Homeobox-leucine zipper protein,
partial (46%)
Length = 701
Score = 95.9 bits (237), Expect(2) = 8e-34
Identities = 57/105 (54%), Positives = 70/105 (66%), Gaps = 10/105 (9%)
Frame = +1
Query: 81 VVSSFSTARVVKRERVDLSCEEIEAEERLSSRV--SDEDD--------DGTNARKKLRLT 130
V S ST + +R + E E SSR SD+DD DG +RKKLRL+
Sbjct: 274 VSSPNSTISSISGKRSEREGNGDENERTSSSRGGGSDDDDGGACGGDGDGEASRKKLRLS 453
Query: 131 KEQSALLEESFKQHSTLNPKQKQALARQLNLRARQVEVWFQNRRA 175
KEQ+ +LEE+FK+H++LNPKQKQALA+QLNL RQVEVWFQNRRA
Sbjct: 454 KEQALVLEETFKEHNSLNPKQKQALAKQLNLMPRQVEVWFQNRRA 588
Score = 65.5 bits (158), Expect(2) = 8e-34
Identities = 29/35 (82%), Positives = 32/35 (90%)
Frame = +3
Query: 176 RTKLKQTEVDCEFLKKCCETLTDENRRLKKELQEL 210
RTKLKQTEVDCE+LK CCE LT+ENRRL KE+QEL
Sbjct: 597 RTKLKQTEVDCEYLKNCCENLTEENRRLHKEVQEL 701
>TC226210 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper protein,
partial (95%)
Length = 1358
Score = 129 bits (325), Expect = 1e-30
Identities = 72/119 (60%), Positives = 84/119 (70%), Gaps = 2/119 (1%)
Frame = +1
Query: 149 PKQKQALAR-QLNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKEL 207
P+QK + + +LRARQVEVWFQNRRARTKLKQTEVDCEFLK+CCE LT ENRRL+KE+
Sbjct: 658 PEQKGKVGSFRQHLRARQVEVWFQNRRARTKLKQTEVDCEFLKRCCENLTVENRRLQKEV 837
Query: 208 QELKALKLAQPLYMPMS-AATLTMCPSCERLGDGGSNIKSPFTITPKPHFFNPFTHPSA 265
QEL+ALKL+ YM M+ TLTMCPSCER+ S+ P H P T P A
Sbjct: 838 QELRALKLSPQFYMHMTPPTTLTMCPSCERVAVPPSSAVDP----AMRHHHVPPTQPRA 1002
Score = 98.6 bits (244), Expect = 3e-21
Identities = 74/197 (37%), Positives = 102/197 (51%), Gaps = 29/197 (14%)
Frame = +2
Query: 9 NPGLHLALGLSLTTTNTSKETTTTTTTSSPKPTVMKPYSSKEPSLTL-GLSGNNKVYCED 67
NP HL L L +++ ++ + KP+ P++S S L G+ N D
Sbjct: 125 NPPTHLHLNLV---------SSSPSSHNPQKPSWNDPFTSSAGSSFLRGIDVNRLPSVVD 277
Query: 68 PLELSRQTSPHSDVVSSFSTARVVKRERVDLSCEEIEAEERLSSRVSDEDDDGTNARKKL 127
E + +SP+S VSS S R ER + + EE + + S + +++D +RKKL
Sbjct: 278 CEEEAGVSSPNS-TVSSVSGKR---SEREEANGEENDTDRACSRGIISDEEDAETSRKKL 445
Query: 128 RLTKEQSALLEESFKQHSTLNP----------------------------KQKQALARQL 159
RL+K+QS +LEESFK+H+TLNP KQK ALA+QL
Sbjct: 446 RLSKDQSIILEESFKEHNTLNPVSN**FV*CPMFLFS*VTNV*ICCVVLQKQKLALAKQL 625
Query: 160 NLRARQVEVWFQNRRAR 176
LRARQVEVWFQNRRAR
Sbjct: 626 GLRARQVEVWFQNRRAR 676
>TC205126 similar to UP|HAT9_ARATH (P46603) Homeobox-leucine zipper protein
HAT9 (Homeodomain-leucine zipper protein HAT9)
(Homeodomain transcription factor HAT9) (HD-ZIP protein
9), partial (25%)
Length = 515
Score = 128 bits (322), Expect = 2e-30
Identities = 61/83 (73%), Positives = 69/83 (82%), Gaps = 5/83 (6%)
Frame = +1
Query: 190 KKCCETLTDENRRLKKELQELKALKLAQPLYMPMSAATLTMCPSCERLG-----DGGSNI 244
++CCETLTDENRRL+KELQELKALKLAQPLYMPM AATL MCPSCERLG G +
Sbjct: 13 EECCETLTDENRRLQKELQELKALKLAQPLYMPMPAATLAMCPSCERLGGSAVNGAGGSP 192
Query: 245 KSPFTITPKPHFFNPFTHPSAAC 267
K+ F++ PKPHFFNPF +PSAAC
Sbjct: 193 KTSFSMAPKPHFFNPFANPSAAC 261
>TC219333 homologue to UP|Q42437 (Q42437) HD-ZIP protein, partial (28%)
Length = 751
Score = 124 bits (310), Expect = 6e-29
Identities = 62/92 (67%), Positives = 70/92 (75%), Gaps = 1/92 (1%)
Frame = +2
Query: 165 QVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS 224
QVEVWFQNRRARTKLKQTEVDCE LK+CCE LT+ENRRL+KE+QEL+ALKL+ YM MS
Sbjct: 2 QVEVWFQNRRARTKLKQTEVDCEVLKRCCENLTEENRRLQKEVQELRALKLSPQFYMQMS 181
Query: 225 -AATLTMCPSCERLGDGGSNIKSPFTITPKPH 255
TLTMCPSCER+ S + S P H
Sbjct: 182 PPTTLTMCPSCERVAVSSSAVGSATRHHPMAH 277
>BG509298 weakly similar to PIR|T09784|T097 homeobox leucine zipper protein
Hb-2 dehydration-inducible - Craterostigma
plantagineum, partial (23%)
Length = 408
Score = 107 bits (266), Expect = 7e-24
Identities = 56/81 (69%), Positives = 65/81 (80%), Gaps = 3/81 (3%)
Frame = +3
Query: 159 LNLRARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLA-Q 217
LNL+ RQVEVWFQNRRARTKLKQTEVD E LKK C+ L+DEN+RLKKELQEL+ALK+
Sbjct: 3 LNLKTRQVEVWFQNRRARTKLKQTEVDHELLKKHCQNLSDENKRLKKELQELRALKVGPS 182
Query: 218 PLYMPMS--AATLTMCPSCER 236
PL + +S A TMC SC+R
Sbjct: 183 PLCIQLSKTATLTTMCSSCDR 245
>BG404990
Length = 362
Score = 105 bits (261), Expect = 3e-23
Identities = 64/108 (59%), Positives = 67/108 (61%), Gaps = 15/108 (13%)
Frame = +1
Query: 1 MGLDHDASNPGLHLALGLSLTTTNTSKETTTTTTTSSP----KPTVMKPYSSKEPSLTLG 56
MGLD DA++ GLHL LGLSL K T KPT KPY SKEPSLTLG
Sbjct: 40 MGLDQDANSSGLHLVLGLSLDWLVLRKPLRPTKPDDHHLCVIKPTPTKPYPSKEPSLTLG 219
Query: 57 LSGN-----------NKVYCEDPLELSRQTSPHSDVVSSFSTARVVKR 93
LSG NKVYCEDPLE SRQTSPHS VVSSFST RV+KR
Sbjct: 220 LSGKGYHVPRNNVAINKVYCEDPLEFSRQTSPHS-VVSSFSTGRVIKR 360
>TC219622 similar to UP|Q6I664 (Q6I664) HD-ZIP protein (Fragment), partial
(64%)
Length = 1015
Score = 97.1 bits (240), Expect = 8e-21
Identities = 48/76 (63%), Positives = 59/76 (77%), Gaps = 1/76 (1%)
Frame = +2
Query: 176 RTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSC 234
RTKLKQTEVDCE+L CCE LT+ENRRL+KE+QEL+ALKL+ LYM M+ TLTMCPSC
Sbjct: 146 RTKLKQTEVDCEYLNICCENLTEENRRLQKEVQELRALKLSPHLYMQMNPPTTLTMCPSC 325
Query: 235 ERLGDGGSNIKSPFTI 250
ER+ ++ S T+
Sbjct: 326 ERVAVSSASSSSSATM 373
>TC215189 similar to UP|Q39941 (Q39941) HAHB-1, partial (70%)
Length = 1315
Score = 94.4 bits (233), Expect = 5e-20
Identities = 63/167 (37%), Positives = 88/167 (51%), Gaps = 6/167 (3%)
Frame = +3
Query: 103 IEAEERLSSRVSDEDDDGTNA-RKKLRLTKEQSALLEESFKQHSTLNPKQKQALARQLNL 161
IE E ++ D DDG+ A KK RL EQ LE+SF+ + L P++K LAR L L
Sbjct: 393 IEHGEEANNAEEDLSDDGSQAGEKKRRLNMEQVKTLEKSFELGNKLEPERKMQLARALGL 572
Query: 162 RARQVEVWFQNRRARTKLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYM 221
+ RQ+ +WFQNRRAR K KQ E D + LK+ E + +N L+ + Q+L+A LA
Sbjct: 573 QPRQIAIWFQNRRARWKTKQLEKDYDVLKRQYEAVKSDNDALQAQNQKLQAEILALKSRE 752
Query: 222 PMSAATLTMCP--SCERLGDGGSNIKSPFTITP---KPHFFNPFTHP 263
P + L SC + S+IK + TP PH + + P
Sbjct: 753 PTESINLNKETEGSCSNRSENSSDIKLDISRTPAIDSPHSTHQQSRP 893
>TC230834 homologue to UP|Q39862 (Q39862) Homeobox-leucine zipper protein,
partial (26%)
Length = 731
Score = 92.8 bits (229), Expect = 1e-19
Identities = 44/61 (72%), Positives = 52/61 (85%), Gaps = 1/61 (1%)
Frame = +3
Query: 178 KLKQTEVDCEFLKKCCETLTDENRRLKKELQELKALKLAQPLYMPMS-AATLTMCPSCER 236
KLKQTEVDCE LK+CCE LT+ENRRL+KE+QEL+ALKL+ YM M+ TLTMCPSCER
Sbjct: 3 KLKQTEVDCEVLKRCCENLTEENRRLQKEVQELRALKLSPQFYMQMTPPTTLTMCPSCER 182
Query: 237 L 237
+
Sbjct: 183 V 185
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.127 0.358
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,194,018
Number of Sequences: 63676
Number of extensions: 142280
Number of successful extensions: 1791
Number of sequences better than 10.0: 156
Number of HSP's better than 10.0 without gapping: 1449
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1653
length of query: 267
length of database: 12,639,632
effective HSP length: 96
effective length of query: 171
effective length of database: 6,526,736
effective search space: 1116071856
effective search space used: 1116071856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Lotus: description of TM0132.1


