

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0083.8
(488 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
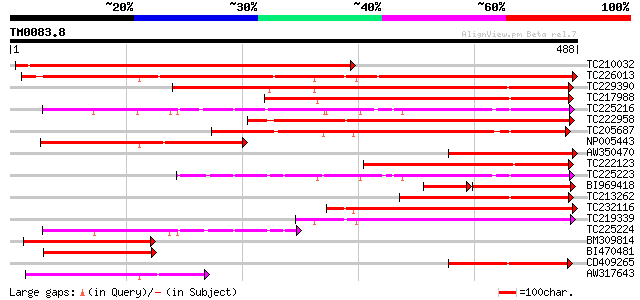
Score E
Sequences producing significant alignments: (bits) Value
TC210032 similar to UP|Q41012 (Q41012) Endo-1,4-beta-glucanase p... 548 e-156
TC226013 similar to UP|Q8L6S6 (Q8L6S6) Endo-beta-1,4-glucanase ... 447 e-126
TC229390 CMCase; cellulase; endo-1,4-beta-D-glucanase 398 e-111
TC217988 similar to UP|Q9SSU7 (Q9SSU7) Endo-1,4-beta-glucanase, ... 334 5e-92
TC225216 homologue to UP|Q6PW04 (Q6PW04) Membrane-anchored endo-... 323 1e-88
TC222958 UP|Q39847 (Q39847) CMCase; cellulase; endo-1,4-beta-D-g... 280 1e-75
TC205687 similar to PIR|E96786|E96786 protein F10A5.13 [imported... 246 2e-65
NP005443 cellulase 224 9e-59
AW350470 homologue to PIR|T06770|T06 cellulase (EC 3.2.1.4) prec... 221 7e-58
TC222123 similar to UP|O81403 (O81403) Endo-1,4-beta-glucanase p... 210 1e-54
TC225223 similar to UP|Q6QLN2 (Q6QLN2) Endo-1,4-beta-glucanase ... 207 1e-53
BI969418 similar to PIR|T06770|T06 cellulase (EC 3.2.1.4) precur... 166 5e-52
TC213262 similar to UP|O22297 (O22297) Acidic cellulase , partia... 191 5e-49
TC232116 similar to PIR|B96527|B96527 protein F27J15.28 [importe... 187 9e-48
TC219339 weakly similar to PIR|T01108|T01108 cellulase T21L14.7... 182 3e-46
TC225224 similar to UP|Q6PW04 (Q6PW04) Membrane-anchored endo-1,... 174 1e-43
BM309814 similar to PIR|E86366|E86 protein F26F24.6 [imported] -... 155 4e-38
BI470481 similar to GP|13383303|db endo-1 4-beta glucanase {Popu... 142 3e-34
CD409265 140 1e-33
AW317643 similar to GP|12484392|gb cellulase {Fragaria x ananass... 134 1e-31
>TC210032 similar to UP|Q41012 (Q41012) Endo-1,4-beta-glucanase precursor ,
partial (58%)
Length = 929
Score = 548 bits (1412), Expect = e-156
Identities = 261/292 (89%), Positives = 278/292 (94%)
Frame = +2
Query: 6 SSFMSLLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFD 65
+S + LLFL+ LLFVGNVQCNPNYREALAKSLLF QGQRSGRLP DQQ+KWRS+SGLFD
Sbjct: 56 NSLVFLLFLLT-LLFVGNVQCNPNYREALAKSLLFFQGQRSGRLPSDQQIKWRSNSGLFD 232
Query: 66 GRLANVDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDY 125
GRLANVDLSGGYYDAGDNVKFNFPMA+TTTMLSW+ IEYGKRMGPQ+KEARAAIRW TDY
Sbjct: 233 GRLANVDLSGGYYDAGDNVKFNFPMAYTTTMLSWATIEYGKRMGPQIKEARAAIRWATDY 412
Query: 126 LLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAA 185
LLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDT RTVYWVSPNNPGSDVAAETAAALAA
Sbjct: 413 LLKCATSTPGRLYVGVGDPNVDHKCWERPEDMDTVRTVYWVSPNNPGSDVAAETAAALAA 592
Query: 186 ASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGA 245
AS+VF+++DPTYS L+RTAQ+VY FALQYQGSYS SLGSAVCPFYCSYSGFKDELLWGA
Sbjct: 593 ASIVFRRLDPTYSNKLLRTAQQVYHFALQYQGSYSHSLGSAVCPFYCSYSGFKDELLWGA 772
Query: 246 AWLFRATNAVKYYNLIKTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKN 297
AWLFRATNAV YYNL+K+LGADDQPDIFSWD+KYAGAHVLLSRRALLNGDKN
Sbjct: 773 AWLFRATNAVSYYNLVKSLGADDQPDIFSWDNKYAGAHVLLSRRALLNGDKN 928
>TC226013 similar to UP|Q8L6S6 (Q8L6S6) Endo-beta-1,4-glucanase , partial
(96%)
Length = 2302
Score = 447 bits (1150), Expect = e-126
Identities = 233/490 (47%), Positives = 316/490 (63%), Gaps = 12/490 (2%)
Frame = +2
Query: 11 LLFLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLAN 70
+L L P F G+ +Y +AL+KSLLF + QRSG LP +Q++ WR+ SGL DG+ +
Sbjct: 152 VLLLCPNFAFAGH-----DYGQALSKSLLFFEAQRSGYLPHNQRVTWRAHSGLQDGKASG 316
Query: 71 VDLSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLK 128
VDL GGYYDAGDNVKF PMAFT TM+SWS IEYGK+M +L A A++WGTDY +K
Sbjct: 317 VDLVGGYYDAGDNVKFGLPMAFTVTMMSWSIIEYGKQMAASGELGHAMEAVKWGTDYFIK 496
Query: 129 CATSTPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASL 188
A P LY VGD N DH CW+RPEDM T R Y + P+NPGSD+A ETAAA+AAAS+
Sbjct: 497 -AHPQPNVLYGEVGDGNTDHYCWQRPEDMTTDRHAYKIDPSNPGSDLAGETAAAMAAASI 673
Query: 189 VFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWL 248
VF++ +P Y+ L+R A +++ FA +Y+G Y S+ A +Y S SG+ DELLW AAWL
Sbjct: 674 VFRRSNPAYAAELLRHAYQLFDFADKYRGKYDSSITVAQ-KYYRSISGYNDELLWAAAWL 850
Query: 249 FRATNAVKYYNLI-----KTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKN-----F 298
++A+N Y + + G F WD KYAG L+++ L+ G F
Sbjct: 851 YQASNNQYYLDYLGRNGDSMGGTGWSMTEFGWDVKYAGVQTLVAK-FLMQGKSGHHAPVF 1027
Query: 299 DQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKK 358
++Y+Q+AE FMC L S + + Q T GGL+F+ +N+Q+VTS +FL T Y+ Y+++
Sbjct: 1028ERYQQKAETFMCSCLGKS-NRNVQKTPGGLIFRQRWNNMQFVTSASFLATVYSDYLASSG 1204
Query: 359 YTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAA 418
C + V P L S+AK QVDY+LG+NP SYMVGYG NFP+R+HHRGSS+ S+
Sbjct: 1205RNLRCSSGNVPPAELLSLAKSQVDYLLGDNPRATSYMVGYGSNFPQRVHHRGSSIVSIKV 1384
Query: 419 HPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVV 478
+P + C GG+ +F S +PN+L GA+VGGP+ D F D+R +Y +EP TY N ++
Sbjct: 1385NPSFVSCRGGYATWFSSKRSDPNLLTGALVGGPDAYDDFADERDNYEQTEPATYNNAPLI 1564
Query: 479 GPLAYFAGIH 488
G LA G H
Sbjct: 1565GILARLGGGH 1594
>TC229390 CMCase; cellulase; endo-1,4-beta-D-glucanase
Length = 1348
Score = 398 bits (1023), Expect = e-111
Identities = 196/351 (55%), Positives = 248/351 (69%), Gaps = 6/351 (1%)
Frame = +3
Query: 141 VGDPNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKL 200
VGDPN DH+CWERPEDMDTART Y V N SDVA ETAAALAAAS+ F+ DP YS+
Sbjct: 84 VGDPNSDHECWERPEDMDTARTTYAVDAPNAASDVAGETAAALAAASMAFRSSDPGYSET 263
Query: 201 LVRTAQKVYQFALQYQGSYSDS--LGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYY 258
L R A +QFA Y+G+YSD+ + CP+YC + G++DELLWGAAWL RAT +
Sbjct: 264 LRRNAITAFQFADNYRGAYSDNDNVKYNACPYYCDFDGYQDELLWGAAWLRRATQDDNFL 443
Query: 259 NLI----KTLGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILP 314
N I KTLGADD + F WD+K+AG +VL+S+ + + + Y+ AE+F+C ++P
Sbjct: 444 NYIQTNGKTLGADDNINEFGWDNKHAGLNVLVSKEVIEGNMYSLESYKSSAESFICTLIP 623
Query: 315 NSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLR 374
SPSS +YT GGL+++ SNLQ+ TSI FL YA Y++ NCGNV V+ TLR
Sbjct: 624 ESPSSHIEYTPGGLVYRPGGSNLQHATSIAFLELVYANYLTHASQAINCGNVYVSAQTLR 803
Query: 375 SVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFH 434
AKRQVDYILG+NP+ +SYMVGY +P+RIHHRGSSLPS+ HPQ I C G +++
Sbjct: 804 QHAKRQVDYILGDNPMGLSYMVGYSNYYPQRIHHRGSSLPSIKDHPQFIACKEG-SIYYN 980
Query: 435 SSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
S+NPNPN+LVGAIVGGP+++D + DDR D+ SEPTTYIN VG LAYFA
Sbjct: 981 STNPNPNVLVGAIVGGPDENDDYVDDRVDFRKSEPTTYINAPFVGVLAYFA 1133
>TC217988 similar to UP|Q9SSU7 (Q9SSU7) Endo-1,4-beta-glucanase, partial
(54%)
Length = 1156
Score = 334 bits (857), Expect = 5e-92
Identities = 158/270 (58%), Positives = 199/270 (73%), Gaps = 4/270 (1%)
Frame = +2
Query: 220 SDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT----LGADDQPDIFSW 275
S++L VCPFYCSYSG++DELLWGAAWL +AT Y N IK LGA + + F W
Sbjct: 2 SNALKPYVCPFYCSYSGYQDELLWGAAWLHKATKNPMYLNYIKVNGQILGAAEFDNTFGW 181
Query: 276 DDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDS 335
D+K+AGA +LLS+ L+ ++ Y+ A+NF+C ++P + SSTQYT GGL+FK+ DS
Sbjct: 182 DNKHAGARILLSKEFLVQRVQSLHDYKGHADNFVCSLIPGTSFSSTQYTPGGLLFKMSDS 361
Query: 336 NLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYM 395
N+QYVTS +FLL TYAKY++ NCG + VTP LR++AK+QVDY+LG+NPL+MSYM
Sbjct: 362 NMQYVTSTSFLLLTYAKYLTQSHMLVNCGGITVTPRRLRTIAKKQVDYLLGDNPLKMSYM 541
Query: 396 VGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSD 455
VGYGP +P+RIHHRGSSLPS+A HP I C GF +S +PNPNILVGAIVGGP++ D
Sbjct: 542 VGYGPRYPRRIHHRGSSLPSIAVHPGKIQCSAGFS-VMNSQSPNPNILVGAIVGGPDEHD 718
Query: 456 GFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
FPD RSDY SEP TYIN +VG LAY A
Sbjct: 719 RFPDQRSDYEQSEPATYINSPLVGALAYLA 808
>TC225216 homologue to UP|Q6PW04 (Q6PW04) Membrane-anchored
endo-1,4-beta-glucanase , complete
Length = 2204
Score = 323 bits (828), Expect = 1e-88
Identities = 202/494 (40%), Positives = 284/494 (56%), Gaps = 36/494 (7%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLAN-----VDLSGGYYDAGDN 83
NY AL K+L+F QRSG+LP + WR +SG+ DG+ ++ DL GGYYDAGD
Sbjct: 314 NYTLALHKALMFFNAQRSGKLPKHNNVSWRGNSGMQDGKSSDHSAAIKDLVGGYYDAGDA 493
Query: 84 VKFNFPMAFTTTMLSWSAIEYGKRM--GPQLKEARAAIRWGTDYLLKCATSTPGRL---- 137
+KFNFP +F+ TMLSWS IEY + +L+ + I+WGTDY LK ST +
Sbjct: 494 IKFNFPASFSITMLSWSVIEYSAKYEAAGELEHVKEIIKWGTDYFLKSFNSTADTITTLA 673
Query: 138 -YVGVGD------PNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVF 190
VG+GD PN DH CW RPEDMD R V + + SD+AAE AAALA+AS+VF
Sbjct: 674 AQVGLGDTSGGDSPN-DHYCWMRPEDMDYDRPV---TECHSCSDLAAEMAAALASASIVF 841
Query: 191 KKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFR 250
K + YSK LV A +++F+ + +G YS GS+ + + + + DE +WG AW++
Sbjct: 842 KD-NKAYSKKLVHGATTLFKFSREQRGRYS--AGSSEASIFYNSTSYWDEYVWGGAWMYF 1012
Query: 251 ATNAVKYYNLIKTLGADDQ-------PD--IFSWDDKYAGAHVLLSR-RALLNGDKNFDQ 300
AT Y L G PD + SWD+K AGA VLLSR R L+ +++
Sbjct: 1013ATGNSSYLKLATAPGLAKHAGAFWGGPDYGVLSWDNKLAGAQVLLSRLRLFLSPGYPYEE 1192
Query: 301 ----YRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSN---LQYVTSITFLLTTYAKY 353
+ + MC LP +S T+GGL+ +L LQYV + FL Y+ Y
Sbjct: 1193ILRTFHNQTSIIMCSYLP--VFTSFNRTKGGLI-QLNHGRPQPLQYVVNAAFLAALYSDY 1363
Query: 354 MSAKKYT-FNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSS 412
+ A + CG + + LRS AK Q+DYILG NP +MSY+VG+G ++PK +HHRG+S
Sbjct: 1364LDAADTPGWYCGPNFFSTDVLRSFAKSQIDYILGNNPRKMSYVVGFGNHYPKHVHHRGAS 1543
Query: 413 LPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTY 472
+P + C GG++ + +S PNPN +VGA+V GP++ DGF D R++Y+++EPT
Sbjct: 1544VPK---NKIKYSCKGGWK-WRDTSKPNPNTIVGAMVAGPDKHDGFHDVRTNYNYTEPTLA 1711
Query: 473 INGAVVGPLAYFAG 486
N +V L +G
Sbjct: 1712GNAGLVAALVALSG 1753
>TC222958 UP|Q39847 (Q39847) CMCase; cellulase; endo-1,4-beta-D-glucanase
(Fragment) , complete
Length = 1045
Score = 280 bits (715), Expect = 1e-75
Identities = 134/284 (47%), Positives = 189/284 (66%), Gaps = 2/284 (0%)
Frame = +3
Query: 205 AQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNL-IKT 263
++ ++ FA +Y+GSYS S CPFYCSYSG++DELLW A+WL++A+ KY + I
Sbjct: 12 SKSLFDFADKYRGSYSGS-----CPFYCSYSGYQDELLWAASWLYKASGESKYLSYSIGN 176
Query: 264 LGADDQPDIFSWDDKYAGAHVLLSRRALLNGDKNFDQYRQEAENFMCKILPNSPSSSTQY 323
G FSWD+KY GA LL+ G K+ +++ + E+F+C ++P S S +
Sbjct: 177 QGWSQAVSEFSWDNKYVGAQTLLTEE-FYGGKKDLAKFKSDVESFICSVMPASSSLQIKT 353
Query: 324 TQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYT-FNCGNVLVTPNTLRSVAKRQVD 382
T GGL+F SNLQY TS T +L ++K ++ +CG+ L TP+ +R+ AK QVD
Sbjct: 354 TPGGLLFTRDSSNLQYATSSTMVLFIFSKILNRNHIDRIHCGSALFTPSQIRAFAKTQVD 533
Query: 383 YILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNI 442
YILG NP++MSYMVG+G +PK++HHRGSS+PS+ HP +GC+ G +++S+NPNPN
Sbjct: 534 YILGSNPMKMSYMVGFGSKYPKQLHHRGSSIPSINVHPTKVGCNDGLSVYYNSANPNPNT 713
Query: 443 LVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAG 486
VGAIVGGP+ +D F D RSDYSHS PTTY+N A V ++ G
Sbjct: 714 HVGAIVGGPDSNDRFSDARSDYSHSGPTTYMNAAFVASVSALLG 845
>TC205687 similar to PIR|E96786|E96786 protein F10A5.13 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(55%)
Length = 1314
Score = 246 bits (628), Expect = 2e-65
Identities = 132/322 (40%), Positives = 197/322 (60%), Gaps = 13/322 (4%)
Frame = +2
Query: 174 DVAAETAAALAAASLVFKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCS 233
D+AAETAAA+A+ASLVFKK DPTYS L++ A++++ FA + +GSYS+++ Y +
Sbjct: 2 DIAAETAAAMASASLVFKKTDPTYSSTLLKHAKQLFTFADKNRGSYSENIPEVAT--YYN 175
Query: 234 YSGFKDELLWGAAWLFRATNAVKYYNLIKTLGADD-----QPDIFSWDDKYAGAHVLLSR 288
+G+ DELLW A+WL+ AT Y + +D P FSWD+K AG VLLSR
Sbjct: 176 STGYGDELLWAASWLYHATGDDSYLQFVTGQDGEDYAQWGSPTWFSWDNKLAGTQVLLSR 355
Query: 289 RALLNG-------DKNFDQYRQEAENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVT 341
+ YR+ AE MC +LP+SP+++ T GL++ ++LQ+
Sbjct: 356 LSFFKAKDISNSYSSGLHSYRKTAEAVMCGLLPDSPTATKSRTDDGLIWVSQWNSLQHPV 535
Query: 342 SITFLLTTYAKYM-SAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGP 400
+ FL Y+ YM +++ CG+ TP+ LR AK Q DY+LG+NP+ MS++VGYG
Sbjct: 536 ASAFLAAVYSDYMLTSQTPKLKCGSDSFTPSDLRDFAKSQADYVLGKNPMHMSFLVGYGD 715
Query: 401 NFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDD 460
+P+ +HHRG+S+P+ A GC GFQ + SS+PNPN+ GA+VGGP ++ F D
Sbjct: 716 KYPQFVHHRGASIPADA----KTGCKDGFQ-WLESSDPNPNVATGALVGGPFLNETFIDS 880
Query: 461 RSDYSHSEPTTYINGAVVGPLA 482
R++ +EP+TY + +VG L+
Sbjct: 881 RNNSMQTEPSTYNSAVIVGLLS 946
>NP005443 cellulase
Length = 653
Score = 224 bits (570), Expect = 9e-59
Identities = 108/180 (60%), Positives = 135/180 (75%), Gaps = 2/180 (1%)
Frame = +1
Query: 27 NPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKF 86
N +Y +AL K++LF +GQRSG LP Q++KWR DS L DG+L NVDL GGYYDAGDNVKF
Sbjct: 109 NYDYGDALGKAILFFEGQRSGNLPATQRVKWRGDSALSDGKLQNVDLIGGYYDAGDNVKF 288
Query: 87 NFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCATSTPGRLYVGVGDP 144
+PMAFTT++LSW+A+EY + QL +AI WG D++L+ TS P LY VGD
Sbjct: 289 GWPMAFTTSLLSWAAVEYESEISSVNQLGYLHSAIHWGADFILRAHTS-PTTLYTQVGDG 465
Query: 145 NVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVRT 204
N DH CWERPEDMDTAR VY + N+PG++ AAE+AAALAAAS+VFKK+D YS L+R+
Sbjct: 466 NADHNCWERPEDMDTARAVYKIDANSPGTEAAAESAAALAAASIVFKKIDANYSSTLLRS 645
>AW350470 homologue to PIR|T06770|T06 cellulase (EC 3.2.1.4) precursor -
garden pea, partial (22%)
Length = 533
Score = 221 bits (562), Expect = 7e-58
Identities = 100/111 (90%), Positives = 104/111 (93%)
Frame = -1
Query: 378 KRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSN 437
KRQVDYILG NPL+MSYMVGYGP FPKR+HHRGSSLPS+ AHPQTIGCDGGF PFFHS N
Sbjct: 533 KRQVDYILGANPLRMSYMVGYGPYFPKRVHHRGSSLPSIEAHPQTIGCDGGFNPFFHSMN 354
Query: 438 PNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAGIH 488
PNPNILVGAIVGGPNQ+DGFPDDRSDYSHSEP TYINGA VGPLAYFAGIH
Sbjct: 353 PNPNILVGAIVGGPNQNDGFPDDRSDYSHSEPATYINGAFVGPLAYFAGIH 201
>TC222123 similar to UP|O81403 (O81403) Endo-1,4-beta-glucanase precursor ,
partial (37%)
Length = 778
Score = 210 bits (535), Expect = 1e-54
Identities = 102/182 (56%), Positives = 128/182 (70%), Gaps = 1/182 (0%)
Frame = +1
Query: 305 AENFMCKILPNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCG 364
A+ F+C LP QY+ GGL+FK SN+Q+VTS++FLL Y+ Y+S CG
Sbjct: 1 ADGFICSTLPGISHPQVQYSPGGLIFKAGGSNMQHVTSLSFLLLAYSNYLSHANKVVPCG 180
Query: 365 NVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIG 424
TP L+ +AKRQVDYILG+NPL MSYMVGYG +P+RIHHR SSLPS+AAHP IG
Sbjct: 181 ETSATPALLKHLAKRQVDYILGDNPLGMSYMVGYGARYPQRIHHRASSLPSVAAHPAHIG 360
Query: 425 CDGGFQPFFHSSNPNPNILVGAIVGGP-NQSDGFPDDRSDYSHSEPTTYINGAVVGPLAY 483
C G + +F S NPNPN+LVGA+VGGP N +D FPD R + SEPTTYIN +VG L++
Sbjct: 361 CKAGSRYYF-SPNPNPNVLVGAVVGGPTNNTDSFPDSRPFFQQSEPTTYINAPLVGLLSF 537
Query: 484 FA 485
F+
Sbjct: 538 FS 543
>TC225223 similar to UP|Q6QLN2 (Q6QLN2) Endo-1,4-beta-glucanase , partial
(61%)
Length = 1470
Score = 207 bits (526), Expect = 1e-53
Identities = 140/362 (38%), Positives = 202/362 (55%), Gaps = 19/362 (5%)
Frame = +3
Query: 144 PNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLVFKKVDPTYSKLLVR 203
PN DH CW RPED+D R + + SD+AAE AAALAAAS+VFK + YS+ LV
Sbjct: 3 PN-DHYCWMRPEDIDYYRPTHTCTTC---SDLAAEMAAALAAASIVFKD-NRAYSQKLVH 167
Query: 204 TAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLFRATNAVKYYNLIKT 263
A +++F+ ++G S + G FY S S + DE +WG AW++ AT Y L T
Sbjct: 168 GATTLFKFSRDHRGRDSPN-GKEASVFYNSTS-YWDEFVWGGAWMYFATGNSSYLKLATT 341
Query: 264 ----------LGADDQPDIFSWDDKYAGAHVLLSR-RALLNGDKNFDQ----YRQEAENF 308
G D + SWD+K GA VLLSR R L+ +++ + +
Sbjct: 342 PRLAKHAGAFWGGPDY-GVLSWDNKLTGAQVLLSRLRLFLSPGYPYEEILTTFHNQTGIV 518
Query: 309 MCKILPNSPSSSTQYTQGGLMFKLPDSN---LQYVTSITFLLTTYAKYMSAKKYT-FNCG 364
MC LP S + T+GGL+ +L LQYV + FL Y+ Y+ A + CG
Sbjct: 519 MCSYLPMFTSFNR--TRGGLI-QLNHGRPQPLQYVVNAAFLAALYSDYLDAADTPGWYCG 689
Query: 365 NVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIG 424
+ + LR AK Q+DYILG+NP +MSY+VG+G ++PK +HHRG+S+P +
Sbjct: 690 PNFFSTDVLRDFAKTQIDYILGKNPRKMSYIVGFGNHYPKHVHHRGASIPK---NKVKYN 860
Query: 425 CDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
C GG++ + SS PNP+ +VGA+V GP++ D F D R++Y+++EPT N +V L
Sbjct: 861 CKGGWK-WRDSSKPNPHTIVGAMVAGPDKHDHFHDVRTNYNYTEPTLAGNAGLVAALVAL 1037
Query: 485 AG 486
+G
Sbjct: 1038SG 1043
>BI969418 similar to PIR|T06770|T06 cellulase (EC 3.2.1.4) precursor - garden
pea, partial (26%)
Length = 510
Score = 166 bits (420), Expect(2) = 5e-52
Identities = 76/89 (85%), Positives = 79/89 (88%)
Frame = -3
Query: 399 GPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFP 458
GP FPKRI H SSLPS+AAHPQ+IGCDGGF PFFHS NPNPNILVGAIVGGPNQ+DGFP
Sbjct: 379 GPYFPKRIRHIXSSLPSIAAHPQSIGCDGGFNPFFHSMNPNPNILVGAIVGGPNQNDGFP 200
Query: 459 DDRSDYSHSEPTTYINGAVVGPLAYFAGI 487
DD SDYSHSEP TYIN A VGPLAYFAGI
Sbjct: 199 DDXSDYSHSEPATYINAAFVGPLAYFAGI 113
Score = 57.0 bits (136), Expect(2) = 5e-52
Identities = 27/41 (65%), Positives = 32/41 (77%)
Frame = -2
Query: 357 KKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVG 397
+K TF VLVTP+ LRS KRQV+YILG NP++MSYMVG
Sbjct: 506 RKXTFXXXKVLVTPSALRSGXKRQVEYILGANPIRMSYMVG 384
>TC213262 similar to UP|O22297 (O22297) Acidic cellulase , partial (30%)
Length = 639
Score = 191 bits (486), Expect = 5e-49
Identities = 95/151 (62%), Positives = 111/151 (72%), Gaps = 1/151 (0%)
Frame = +1
Query: 336 NLQYVTSITFLLTTYAKYMSAKK-YTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSY 394
NLQYVTS +FLL TYAKY++ CG VT L ++AK QVDYILG NP +MSY
Sbjct: 4 NLQYVTSTSFLLLTYAKYLNTNGGNVVRCGTSAVTGENLVTLAKAQVDYILGNNPTKMSY 183
Query: 395 MVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQS 454
MVG+G +PK IHHRGSSLPS+ AH Q I C+ GFQ FFHS++PNPNILVGAIVGGP+ +
Sbjct: 184 MVGFGERYPKHIHHRGSSLPSIHAHTQHISCNDGFQ-FFHSASPNPNILVGAIVGGPDNN 360
Query: 455 DGFPDDRSDYSHSEPTTYINGAVVGPLAYFA 485
D F DDR +Y SEP TYIN VG LAYF+
Sbjct: 361 DNFSDDRHNYQQSEPATYINAPFVGALAYFS 453
>TC232116 similar to PIR|B96527|B96527 protein F27J15.28 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(50%)
Length = 1248
Score = 187 bits (475), Expect = 9e-48
Identities = 91/221 (41%), Positives = 134/221 (60%), Gaps = 5/221 (2%)
Frame = +3
Query: 273 FSWDDKYAGAHVLLSRRALLNG-----DKNFDQYRQEAENFMCKILPNSPSSSTQYTQGG 327
FSWD+KYAG +LLS+ LL G QY+ +AE F C L + + Q T GG
Sbjct: 63 FSWDNKYAGVQILLSK-VLLEGKAGAYSATLKQYQAKAEYFTCACLQKNDDYNVQKTPGG 239
Query: 328 LMFKLPDSNLQYVTSITFLLTTYAKYMSAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGE 387
L++ +N+QYV+S FLL Y+ Y+SA K NC + P L + K Q DYILG+
Sbjct: 240 LLYVREWNNMQYVSSAAFLLAVYSNYLSATKSQLNCPDGQTQPQELLNFVKSQADYILGK 419
Query: 388 NPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAI 447
NP +SY+VGYG +P +HHRG+S+ S+ A +GC GF+ +++ + PNPN++ G +
Sbjct: 420 NPADVSYLVGYGAKYPLHVHHRGASIASIFALHYDVGCTQGFETWYNRAEPNPNVICGGL 599
Query: 448 VGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYFAGIH 488
VGGP+++D F D+RS+Y +EPT + +VG A ++
Sbjct: 600 VGGPDKNDDFSDERSNYEQTEPTISGSAPLVGIFAKLQSLY 722
>TC219339 weakly similar to PIR|T01108|T01108 cellulase T21L14.7 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(49%)
Length = 881
Score = 182 bits (462), Expect = 3e-46
Identities = 97/253 (38%), Positives = 148/253 (58%), Gaps = 12/253 (4%)
Frame = +3
Query: 247 WLFRATNAVKYYNLI----KTLGADDQP-DIFSWDDKYAGAHVLLSRRALLNGDKN---- 297
WL++AT+ Y + T G F WD KYAG +++S+ LL+ +K+
Sbjct: 6 WLYKATDNKMYLQYVISNAHTFGGTGWSISEFIWDVKYAGLQLMVSK--LLSEEKHKKHR 179
Query: 298 --FDQYRQEAENFMCKIL-PNSPSSSTQYTQGGLMFKLPDSNLQYVTSITFLLTTYAKYM 354
+QY+ +AE ++C L N+ S++ + T GL++ +N+QYV++ FLL+ Y+ ++
Sbjct: 180 DILEQYKSKAEYYICSCLNKNNDSNNVERTPAGLIYVRQWNNMQYVSTAAFLLSIYSDFL 359
Query: 355 SAKKYTFNCGNVLVTPNTLRSVAKRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLP 414
+ NC V + + AK Q DYILG NP+ MSY+VGYGPN+PKR+HHRG+S+
Sbjct: 360 QSTNQKLNCHGGTVDHEEILNFAKSQADYILGSNPMNMSYLVGYGPNYPKRVHHRGASIV 539
Query: 415 SLAAHPQTIGCDGGFQPFFHSSNPNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYIN 474
S + IGC G+ ++ S PNPN+LVGA+VGGP+ D F D R+++ +E TY
Sbjct: 540 SYKKNKGFIGCTQGYDNWYGSQAPNPNVLVGALVGGPDGKDNFEDRRNNFMQTEACTYNT 719
Query: 475 GAVVGPLAYFAGI 487
+VG A F I
Sbjct: 720 APLVGVFAKFLHI 758
>TC225224 similar to UP|Q6PW04 (Q6PW04) Membrane-anchored
endo-1,4-beta-glucanase , partial (54%)
Length = 1078
Score = 174 bits (440), Expect = 1e-43
Identities = 113/242 (46%), Positives = 143/242 (58%), Gaps = 19/242 (7%)
Frame = +2
Query: 29 NYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANV-----DLSGGYYDAGDN 83
NY AL K+LLF Q+SG+LP + WR +SGL DG+ V DL GGYYDAGD
Sbjct: 374 NYTIALHKALLFFNAQKSGKLPRHNNVSWRGNSGLQDGKGDGVPPVIKDLVGGYYDAGDA 553
Query: 84 VKFNFPMAFTTTMLSWSAIEY-GK-RMGPQLKEARAAIRWGTDYLLKCATSTPGR----- 136
+KFNFPM+F+ TMLSWS IEY GK + +L + I+WGTDYLLK ST
Sbjct: 554 IKFNFPMSFSMTMLSWSVIEYSGKYQAAGELGHVKDIIKWGTDYLLKNFNSTADTITQLG 733
Query: 137 LYVGVGD-------PNVDHKCWERPEDMDTARTVYWVSPNNPGSDVAAETAAALAAASLV 189
+ VG GD PN DH CW RPED+D R + SD+AAE AAALAAAS+V
Sbjct: 734 MQVGSGDTSQGSTTPN-DHYCWMRPEDIDYDRPTQTCT---TCSDLAAEMAAALAAASIV 901
Query: 190 FKKVDPTYSKLLVRTAQKVYQFALQYQGSYSDSLGSAVCPFYCSYSGFKDELLWGAAWLF 249
FK + YS+ LV A +++F+ +G YS + G FY S S + DE +WG A ++
Sbjct: 902 FKD-NRAYSQKLVHGATTLFKFSRDQRGRYSPN-GKEASVFYNSTS-YWDEFVWGGAXMY 1072
Query: 250 RA 251
A
Sbjct: 1073XA 1078
>BM309814 similar to PIR|E86366|E86 protein F26F24.6 [imported] - Arabidopsis
thaliana, partial (22%)
Length = 398
Score = 155 bits (392), Expect = 4e-38
Identities = 72/113 (63%), Positives = 86/113 (75%)
Frame = +3
Query: 13 FLIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVD 72
FL +LL + + NYR+AL KS+LF +GQRSG+LP DQ+L+WR DS L DG A VD
Sbjct: 60 FLAMLLLLLRSTAAGHNYRDALRKSILFFEGQRSGKLPADQRLRWRHDSALHDGATAGVD 239
Query: 73 LSGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDY 125
LSGGYYDAGDN+KF FPMAFTTTMLSWS I++ K MG +L A A+RWGTDY
Sbjct: 240 LSGGYYDAGDNIKFGFPMAFTTTMLSWSVIDFEKSMGAELGNALKAVRWGTDY 398
>BI470481 similar to GP|13383303|db endo-1 4-beta glucanase {Populus alba},
partial (24%)
Length = 415
Score = 142 bits (358), Expect = 3e-34
Identities = 67/97 (69%), Positives = 76/97 (78%)
Frame = +2
Query: 30 YREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDLSGGYYDAGDNVKFNFP 89
Y EAL KS+LF +GQRSG+LP +QQ WR DSGL DG +VDL GGYYDAGDNVKF P
Sbjct: 125 YHEALEKSILFFEGQRSGKLPSNQQQTWRGDSGLSDGSSYHVDLVGGYYDAGDNVKFGLP 304
Query: 90 MAFTTTMLSWSAIEYGKRMGPQLKEARAAIRWGTDYL 126
MAFTTT+L+WS IE+G M QL+ ARAAIRW TDYL
Sbjct: 305 MAFTTTLLAWSVIEFGSSMQDQLENARAAIRWSTDYL 415
>CD409265
Length = 681
Score = 140 bits (354), Expect = 1e-33
Identities = 67/107 (62%), Positives = 81/107 (75%)
Frame = -1
Query: 378 KRQVDYILGENPLQMSYMVGYGPNFPKRIHHRGSSLPSLAAHPQTIGCDGGFQPFFHSSN 437
K+QVDYILG+NP+ MSYMVGY +P+ IHHRGSSLPS+ HPQ I C G +F+SS+
Sbjct: 681 KKQVDYILGDNPMGMSYMVGYSYKYPQHIHHRGSSLPSIKDHPQFIACKEG-SIYFNSSH 505
Query: 438 PNPNILVGAIVGGPNQSDGFPDDRSDYSHSEPTTYINGAVVGPLAYF 484
P P +LVGAIVGGP + D + DDR+D+ SEPTTYIN VG LAYF
Sbjct: 504 PYPYVLVGAIVGGPGEDDVYEDDRADFRKSEPTTYINAPFVGILAYF 364
>AW317643 similar to GP|12484392|gb cellulase {Fragaria x ananassa}, partial
(25%)
Length = 511
Score = 134 bits (336), Expect = 1e-31
Identities = 76/161 (47%), Positives = 95/161 (58%), Gaps = 2/161 (1%)
Frame = +2
Query: 14 LIPMLLFVGNVQCNPNYREALAKSLLFLQGQRSGRLPPDQQLKWRSDSGLFDGRLANVDL 73
L+ +LL +Y +AL+ SL F + RSG L Q + WR+ SGL DG+ + VDL
Sbjct: 29 LLALLLCANCAFAGHDYGQALSTSLQFFETHRSGCLSHTQMVTWRAHSGLQDGKSSGVDL 208
Query: 74 SGGYYDAGDNVKFNFPMAFTTTMLSWSAIEYGKRMGP--QLKEARAAIRWGTDYLLKCAT 131
G YYDA DNVKF PMAFT TM+SWS IEYG++M +L A A + GTD + A
Sbjct: 209 LGRYYDARDNVKFGLPMAFTDTMMSWSIIEYGQQMAASGELGHALEASK*GTDCFI-IAH 385
Query: 132 STPGRLYVGVGDPNVDHKCWERPEDMDTARTVYWVSPNNPG 172
+ LY V D N DH CW+RP+DM T R Y V P N G
Sbjct: 386 TQANVLYREVRDGNTDHCCWQRPQDMTTDRHAYKVHPINSG 508
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,483,980
Number of Sequences: 63676
Number of extensions: 355715
Number of successful extensions: 1604
Number of sequences better than 10.0: 60
Number of HSP's better than 10.0 without gapping: 1543
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1551
length of query: 488
length of database: 12,639,632
effective HSP length: 101
effective length of query: 387
effective length of database: 6,208,356
effective search space: 2402633772
effective search space used: 2402633772
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Lotus: description of TM0083.8


