

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0070.9
(437 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
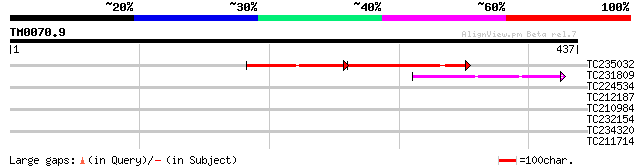
TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarit... 94 9e-37
TC231809 42 4e-04
TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), p... 40 0.002
TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragm... 37 0.013
TC210984 37 0.023
TC232154 weakly similar to GB|AAF86996.1|9295690|AC005292 F26F24... 31 1.3
TC234320 29 3.7
TC211714 weakly similar to UP|Q94KC2 (Q94KC2) Polyphenol oxidase... 29 4.8
>TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarity to
ribosomal protein, partial (24%)
Length = 689
Score = 94.4 bits (233), Expect(2) = 9e-37
Identities = 49/79 (62%), Positives = 57/79 (72%)
Frame = +3
Query: 183 LLQWGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHA 242
L+Q E RGF MLGSID +HW+ K C VAWKGQY GD G PTIML+ V SQ L I HA
Sbjct: 21 LMQICEIRGFQSMLGSIDYLHWK*KTCLVAWKGQYCHGD-GPPTIMLKVVVSQYLRI*HA 197
Query: 243 FFGIAGSNNDITVLNQSPV 261
+ G+ GSNN+I VLN+S V
Sbjct: 198 YLGVEGSNNNINVLNESCV 254
Score = 77.8 bits (190), Expect(2) = 9e-37
Identities = 42/95 (44%), Positives = 58/95 (60%)
Frame = +1
Query: 261 VFNEVLRGAAPMVKFRVNETMYHIGYYLADGIYPEWGTFVKTIPMPQGEKKQKFAKRQEA 320
VF +V+ G AP+V+F VNE Y++GY+L GIY +W T VKTI P G K++ F + QEA
Sbjct: 250 VFKDVMEGEAPIVQFAVNEIQYNMGYFLTYGIYLDWTTLVKTILRP*GHKRKLFTQYQEA 429
Query: 321 ARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSII 355
RKD + + FAI+RG W N ++ I
Sbjct: 430 TRKDRI*STSIW---FAIIRGLVCGWSMNTYQAYI 525
Score = 38.1 bits (87), Expect = 0.008
Identities = 22/61 (36%), Positives = 32/61 (52%), Gaps = 4/61 (6%)
Frame = +3
Query: 345 WWHPNDMKSI--IYACIILHNMIVEDERNTYKGNF--VYEQVNNDISDAEVLSGPIPAFR 400
W + SI +Y CIILHNMIV+DE +TY N Y+ N++ ++ G P
Sbjct: 492 WLEHEHISSIYYMYVCIILHNMIVKDEPHTYNENIDRQYKYPTNNVLIIDIHIGLHPVLN 671
Query: 401 N 401
+
Sbjct: 672 H 674
>TC231809
Length = 760
Score = 42.4 bits (98), Expect = 4e-04
Identities = 34/118 (28%), Positives = 55/118 (45%)
Frame = +3
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDER 370
K+ F R R + R+F VL++RF I++ ++ +I C+ LHN I +ER
Sbjct: 207 KELFNHRHCFLRNAILRSFNVLKTRFPILKLAPQYSFQIQRDIVIAGCV-LHNFIRREER 383
Query: 371 NTYKGNFVYEQVNNDISDAEVLSGPIPAFRNMLERRAHQIEKSIHRQLQADLVEHIWD 428
N + N V +ISD + L +P ++ E+ A SI + D + WD
Sbjct: 384 NDWIFNSVGGSPVEEISDIDELP-DVPMLSSIQEQLAFSSRDSIASSMWDDFLTK-WD 551
>TC224534 similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment), partial (50%)
Length = 968
Score = 40.4 bits (93), Expect = 0.002
Identities = 23/64 (35%), Positives = 41/64 (63%), Gaps = 3/64 (4%)
Frame = +1
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKS---IIYACIILHNMIVE 367
K+ F R +A+ +ER+FGVL+ R++I+R PS + D+K+ II C +L N I +
Sbjct: 589 KELFNLRHASAKNVIERSFGVLKKRWSILRTPSFF----DIKTQIRIINTCFMLRNFIRD 756
Query: 368 DERN 371
++++
Sbjct: 757 EQQS 768
>TC212187 weakly similar to UP|Q6USA0 (Q6USA0) Transposase (Fragment),
partial (45%)
Length = 973
Score = 37.4 bits (85), Expect = 0.013
Identities = 31/119 (26%), Positives = 56/119 (47%), Gaps = 12/119 (10%)
Frame = +1
Query: 284 IGYYLADGIYPEW----GTFVKT-IPMPQGEKKQKFAKRQE-------AARKDVERAFGV 331
+ +YL D YP + G++ KT +PQ + R E + + +ERAFG+
Sbjct: 508 VNFYLVDSCYPTFMGFLGSYNKTRYHLPQFRIGPRIRGRVEVFNYYHSSLQSTIERAFGL 687
Query: 332 LQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDERNTYKGNFVYEQVNNDISDAE 390
++R+ I+ S + II AC+ +HN I ++++ G F +N+ D +
Sbjct: 688 CKARWKILGNMSPF-ALKTQNQIIVACMTIHNFIQRNDKS--DGEFDSLDEDNEYIDTD 855
>TC210984
Length = 1085
Score = 36.6 bits (83), Expect = 0.023
Identities = 23/82 (28%), Positives = 39/82 (47%)
Frame = +3
Query: 311 KQKFAKRQEAARKDVERAFGVLQSRFAIVRGPSRWWHPNDMKSIIYACIILHNMIVEDER 370
K+ F R + R +ER FG+ +SRF I + + + ++ AC LHN + ++ R
Sbjct: 423 KELFNLRHASLRNVIERIFGIFKSRFTIFKSAPPFLFKTQAE-LVLACAALHNFLRKECR 599
Query: 371 NTYKGNFVYEQVNNDISDAEVL 392
+ F E + S + VL
Sbjct: 600 S---DEFPVEPTDESSSSSSVL 656
>TC232154 weakly similar to GB|AAF86996.1|9295690|AC005292 F26F24.7
{Arabidopsis thaliana;} , partial (40%)
Length = 576
Score = 30.8 bits (68), Expect = 1.3
Identities = 14/46 (30%), Positives = 21/46 (45%)
Frame = -1
Query: 204 WEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWHAFFGIAGS 249
W+W+ C AW + HG ++L + W+W AF GS
Sbjct: 150 WDWEWCACAWPCYASLWSHGSALLVLVVLP----WLWKAFERKEGS 25
>TC234320
Length = 739
Score = 29.3 bits (64), Expect = 3.7
Identities = 15/56 (26%), Positives = 25/56 (43%)
Frame = +1
Query: 186 WGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGDHGRPTIMLEAVASQDLWIWH 241
WG RG+ C+ +NC WK + + +HG +I V + ++I H
Sbjct: 268 WGSGRGY--------CVR---RNCCYDWKALFRKKNHGAWSIRCRKVGREKMFILH 402
>TC211714 weakly similar to UP|Q94KC2 (Q94KC2) Polyphenol oxidase, partial
(24%)
Length = 458
Score = 28.9 bits (63), Expect = 4.8
Identities = 15/49 (30%), Positives = 21/49 (42%)
Frame = +3
Query: 173 RRPNEEDMTRLLQWGESRGFPGMLGSIDCMHWEWKNCPVAWKGQYTRGD 221
R+PN+EDM R G F ++D M WK P + + D
Sbjct: 150 RQPNKEDMGRFYSAGRDPAFYSHHANVDRMWNIWKTIPSGKRRDFKNRD 296
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,704,327
Number of Sequences: 63676
Number of extensions: 250584
Number of successful extensions: 1343
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1333
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1341
length of query: 437
length of database: 12,639,632
effective HSP length: 100
effective length of query: 337
effective length of database: 6,272,032
effective search space: 2113674784
effective search space used: 2113674784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Lotus: description of TM0070.9


