

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0069.5
(595 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
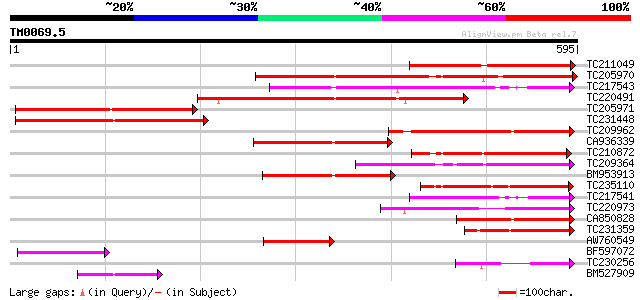
Score E
Sequences producing significant alignments: (bits) Value
TC211049 similar to UP|Q9LQV3 (Q9LQV3) F10B6.18, partial (19%) 280 1e-75
TC205970 278 4e-75
TC217543 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, parti... 238 4e-63
TC220491 223 2e-58
TC205971 199 4e-51
TC231448 169 4e-42
TC209962 153 2e-37
CA936339 149 3e-36
TC210872 146 2e-35
TC209364 144 1e-34
BM953913 138 8e-33
TC235110 similar to PIR|C86410|C86410 protein F3M18.18 [imported... 122 3e-28
TC217541 115 4e-26
TC220973 111 1e-24
CA850828 weakly similar to GP|14209545|dbj P0481E12.26 {Oryza sa... 109 4e-24
TC231359 96 6e-20
AW760549 86 6e-17
BF597072 73 3e-13
TC230256 73 4e-13
BM527909 weakly similar to PIR|C86410|C864 protein F3M18.18 [imp... 72 7e-13
>TC211049 similar to UP|Q9LQV3 (Q9LQV3) F10B6.18, partial (19%)
Length = 668
Score = 280 bits (717), Expect = 1e-75
Identities = 140/175 (80%), Positives = 151/175 (86%), Gaps = 1/175 (0%)
Frame = +1
Query: 420 DDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFS 479
D RF EAI+GKKF HVCTAPVKY+P WSSDKDVAFIVTG QLHVKKHESRSVLHLRLLFS
Sbjct: 1 DGRFFEAINGKKFYHVCTAPVKYDPRWSSDKDVAFIVTGAQLHVKKHESRSVLHLRLLFS 180
Query: 480 KVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQ-KKPALVVVDSSVFPTG 538
KVSN VVKS+WTQGS SG S +FS ISTSI GKDQ +K +V++DSSVFPTG
Sbjct: 181 KVSNCAVVKSSWTQGS-----SGLSQRSGIFSVISTSISGKDQNQKKPVVLLDSSVFPTG 345
Query: 539 PPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLNT 593
PPVPVQ QKLLKF+DTSQLCKGPQDSPGHWL+TGARL+LDK KICLWAKFSLLNT
Sbjct: 346 PPVPVQTQKLLKFIDTSQLCKGPQDSPGHWLITGARLVLDKRKICLWAKFSLLNT 510
>TC205970
Length = 1317
Score = 278 bits (712), Expect = 4e-75
Identities = 156/340 (45%), Positives = 212/340 (61%), Gaps = 3/340 (0%)
Frame = +3
Query: 259 GGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDL 318
G + +HSEW TV ++PD + SF+P TSLL G G+L+HA+NLYLRYKP + +L
Sbjct: 3 GNGKRSLSHSEWCQTVLSQPDVISMSFVPITSLLGGINGSGYLTHAMNLYLRYKPQIEEL 182
Query: 319 PYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGM 378
FL++Q R WAP+ +L LGP + ++ SL + MGPKLYVNT V VGK+PVTG+
Sbjct: 183 HQFLEFQLPRQWAPVFGELALGP-ERKPQNTASLQFSFMGPKLYVNTTPVDVGKKPVTGL 359
Query: 379 RLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKKFSHVCTA 438
RL+LEG + N LAIHLQ+L + P + E S + + ++ E + K FSHVCTA
Sbjct: 360 RLYLEGKRSNCLAIHLQHLSSLPKTFQLQDEPNGNASNDSSERKYYEKVQWKSFSHVCTA 539
Query: 439 PVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGS--- 495
PV SD D A +VTG V + VL LRL F KV A VK +GS
Sbjct: 540 PV------DSDDDNA-VVTGAHFEVGDTGLKKVLFLRLHFCKVVGATRVKVPEWEGSPGL 698
Query: 496 TQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTS 555
TQ+SG S S+ FS Q +P+ V ++S+++P GPPVP Q+ KLL+FVDT+
Sbjct: 699 TQKSGIISTLISTTFSGP----QKPPPPRPSDVNINSALYPGGPPVPTQSPKLLRFVDTT 866
Query: 556 QLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLNTSP 595
++ +GPQDSPG+W+V+GARL+++K KI L K+SLL P
Sbjct: 867 EMTRGPQDSPGYWVVSGARLLVEKAKISLKVKYSLLTVIP 986
>TC217543 similar to UP|Q940R0 (Q940R0) At1g29690/F15D2_24, partial (29%)
Length = 1093
Score = 238 bits (608), Expect = 4e-63
Identities = 144/327 (44%), Positives = 191/327 (58%), Gaps = 7/327 (2%)
Frame = +1
Query: 273 TVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQAHRLWAP 332
TV PD ++ +F P SLL+ PG +L+ AI+LYL YKPP+ DL YFLD+Q R+WAP
Sbjct: 7 TVKLAPDIINMNFTPIVSLLEGVPGVKYLARAIDLYLEYKPPIEDLQYFLDFQITRVWAP 186
Query: 333 IHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGMKCNRLAI 392
N+L + SL +LMGPKL+V+ +VTVG++PVTG+RL LEG K NRLAI
Sbjct: 187 EQNNL-----QRKEPVCQSLQFSLMGPKLFVSPDQVTVGRKPVTGLRLSLEGSKQNRLAI 351
Query: 393 HLQYLLNTPTMLN-----NKIEDTTTW-SEEIIDDRFLEAISGKKFSHVCTAPVKYNPSW 446
HLQ+L++ P L + W E D R+ E I K FSHV TAP++Y +
Sbjct: 352 HLQHLVSLPKNLQPHWDAHMAIGAPKWHGPEEQDSRWFEPIKWKNFSHVSTAPIEYTETS 531
Query: 447 SSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQG-STQRSGSGSGS 505
D IVTG QL V +++VLHL+LLFSKV + +S W ST + G+
Sbjct: 532 IGDLSGVHIVTGAQLGVWDFGAKNVLHLKLLFSKVPGCTIRRSVWDHNPSTPAAQRSDGA 711
Query: 506 GSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSP 565
SS+ T +D+K+ DSS + KL K VD +++ KGPQD P
Sbjct: 712 SSSL-----TKKTSEDKKE------DSS---------IHIGKLAKIVDMTEMSKGPQDIP 831
Query: 566 GHWLVTGARLMLDKGKICLWAKFSLLN 592
GHWLVTGA+L ++KGKI L K+SLLN
Sbjct: 832 GHWLVTGAKLGVEKGKIVLRIKYSLLN 912
>TC220491
Length = 875
Score = 223 bits (568), Expect = 2e-58
Identities = 129/294 (43%), Positives = 182/294 (61%), Gaps = 10/294 (3%)
Frame = +2
Query: 198 KNLDELGDQLFTGTCNFLPK----TKDQKHKVPPAFD-VFGPQIVAFNNSTCVCAKDGIT 252
++L+ LGD LF+ + + T + K KVP F+ V F + + +KDG+T
Sbjct: 2 RHLENLGDFLFSDLKSPSQQQRNSTPEGKQKVPEVFNRVMQSSTTQFASISEASSKDGLT 181
Query: 253 VICAKRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYK 312
+IC+KRGGD +HS WL TV + P+A+ F F+P +SLL PG G+LSHAINLYLRYK
Sbjct: 182 IICSKRGGDVFKQSHSNWLQTVASNPEAILFKFVPISSLLTGIPGSGYLSHAINLYLRYK 361
Query: 313 PPLSDLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGK 372
PP DL FL++Q R WAP+ DLPL + SPSL M PKL+V+ A+V +
Sbjct: 362 PPPDDLQCFLEFQIPRQWAPMFYDLPLRH-QRKKCSSPSLQFGFMFPKLHVSCAQVVSEQ 538
Query: 373 RPVTGMRLFLEGMKCNRLAIHLQYLLNTP-TMLNNKIEDTTTW---SEEIIDDRFLEAIS 428
+PV G+RL+LEG K +RLAIH+ +L + P TM+ + T++W + D FLE I
Sbjct: 539 KPVVGLRLYLEGRKSDRLAIHVHHLSSLPNTMIYS--SGTSSWRGSDDNESSDIFLEPIR 712
Query: 429 GKKFSHVCTAPVKYNPSWSSD-KDVAFIVTGVQLHVKKHESRSVLHLRLLFSKV 481
K F++VCTA VK++P+W + +IVTG QL K ++VLHLRLL++ +
Sbjct: 713 WKGFANVCTAVVKHDPTWLQETSGGVYIVTGAQLISKGTWPKNVLHLRLLYTHI 874
>TC205971
Length = 673
Score = 199 bits (505), Expect = 4e-51
Identities = 95/191 (49%), Positives = 131/191 (67%)
Frame = +1
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKGD 66
E A+ ++G G+DLT+D +LKFCK RL+ +++ R + +P + +V IKCDKGD
Sbjct: 103 EDAIRAIGLGYDLTNDLKLKFCKNHSRLIAIDDDNLRTVELPPRISIPNVPKSIKCDKGD 282
Query: 67 LTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYV 126
R SD+LSF QMSE FNQ S+ GKIP+G+FNT FGF G W DAANTK L DG
Sbjct: 283 RMRLCSDVLSFQQMSEQFNQDLSLSGKIPTGHFNTAFGF-TGVWQKDAANTKTLAFDGVS 459
Query: 127 IKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQD 186
I L+++ ++ +VL V +AVPSSWDP AL RFIEK+GTH++VG+ +GG D++ KQ
Sbjct: 460 ITLYDIAFEKTQVVLHDHVKQAVPSSWDPAALTRFIEKYGTHVIVGVKMGGTDIIYAKQQ 639
Query: 187 VSSNLEPSELK 197
SS + P+E++
Sbjct: 640 YSSTVPPAEVQ 672
>TC231448
Length = 813
Score = 169 bits (427), Expect = 4e-42
Identities = 84/202 (41%), Positives = 129/202 (63%)
Frame = +3
Query: 7 EKALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKGD 66
+ A+NS+G GFD+T D CK RL+ +NE + R L +PG + +V IKC +G+
Sbjct: 108 QNAINSIGLGFDITQDISFDNCKKGSRLIFVNEEQCRHLEIPGGVSIPNVPNSIKCVRGE 287
Query: 67 LTRYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYV 126
R++SD+L+ QM E FNQ+ + G + SG+ FG S D A+ K L DG+
Sbjct: 288 SIRFESDVLTRDQMMEHFNQQMLLSGNLASGHLCASFGLSDRS-IKDLASIKSLAYDGWF 464
Query: 127 IKLFNVHIDRYPLVLSKQVIEAVPSSWDPTALARFIEKFGTHILVGLSIGGKDLVLVKQD 186
IK + + ++R+ + QV EAVPSSWDP ALARFI++FGTH++VG+S+GGKD++ ++Q+
Sbjct: 465 IKRYTIELERHHCKILDQVEEAVPSSWDPEALARFIQRFGTHVIVGVSMGGKDVLYLRQE 644
Query: 187 VSSNLEPSELKKNLDELGDQLF 208
+S L P+ ++K L + + F
Sbjct: 645 DTSYLGPTSIQKLLKDTASRKF 710
>TC209962
Length = 659
Score = 153 bits (386), Expect = 2e-37
Identities = 86/196 (43%), Positives = 122/196 (61%), Gaps = 1/196 (0%)
Frame = +3
Query: 398 LNTPTMLNNKIEDTTTWSEEIIDDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVT 457
L+TP+M ++ + D+FLE I K+FS+VCTA VK++P+W ++ +IVT
Sbjct: 3 LSTPSMWRGSDDNESR-------DQFLERIRWKRFSNVCTAVVKHDPNWLNNS*GVYIVT 161
Query: 458 GVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQG-STQRSGSGSGSGSSVFSAISTS 516
G QL K R+VLHLRLLF+ N + KS WT R S + S+ FS
Sbjct: 162 GAQLLSKGSWPRNVLHLRLLFAHTPNCSIRKSEWTAAPEASRKSSFLTNLSTTFSFTQHG 341
Query: 517 IQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLM 576
G ++ P V++S V+P GPPVPV+A KLLK+V+T+++ +GP D+PGHWLVT A+L+
Sbjct: 342 NTGPPKQAPT--VLNSGVYPDGPPVPVRAGKLLKYVETAEVVRGPHDAPGHWLVTAAKLV 515
Query: 577 LDKGKICLWAKFSLLN 592
D GKI L KF+LL+
Sbjct: 516 TDGGKIGLQVKFALLD 563
>CA936339
Length = 431
Score = 149 bits (377), Expect = 3e-36
Identities = 70/145 (48%), Positives = 97/145 (66%)
Frame = +1
Query: 257 KRGGDTQVSNHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLS 316
+RGGD +HS WL TV + PD + +F P T L+ PG+ L+HAI LYL YKPP+
Sbjct: 1 RRGGDDLEQDHSMWLRTVWSSPDVIQMTFCPITDLIDEVPGKEQLTHAIGLYLEYKPPIE 180
Query: 317 DLPYFLDYQAHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVT 376
+L YFL++Q +WAP+H +P + PSL ++MG KLYV+ ++TVG+ PVT
Sbjct: 181 ELRYFLEFQIAHVWAPLHERIP--GQQRKEPICPSLQFSIMGQKLYVSQEQITVGRLPVT 354
Query: 377 GMRLFLEGMKCNRLAIHLQYLLNTP 401
G+RLFLEG K NRL++HLQ+L + P
Sbjct: 355 GLRLFLEGSKQNRLSVHLQHLSSLP 429
>TC210872
Length = 484
Score = 146 bits (369), Expect = 2e-35
Identities = 82/168 (48%), Positives = 116/168 (68%)
Frame = +3
Query: 422 RFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKV 481
RF E + K FSHVCTAPV+ S++D++ IVTG QL V+ + +++L LRL FS V
Sbjct: 3 RFYEKVQWKNFSHVCTAPVE------SEEDLS-IVTGAQLQVENYGIKNILFLRLRFSTV 161
Query: 482 SNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPV 541
A VK +GS + G+ SG S++ S TS K +PA V ++S+V+P GPPV
Sbjct: 162 LGAKAVKHPEWEGSL-KLGAKSGLISTLISQHFTSTFQKPPPRPADVNINSAVYPGGPPV 338
Query: 542 PVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFS 589
PVQA KLLKFVDT+++ +GPQ+SPG+W+++GA+L++DKGKI L K+S
Sbjct: 339 PVQAPKLLKFVDTTEMTRGPQESPGYWVISGAKLVVDKGKISLRVKYS 482
>TC209364
Length = 725
Score = 144 bits (362), Expect = 1e-34
Identities = 89/230 (38%), Positives = 124/230 (53%), Gaps = 1/230 (0%)
Frame = +2
Query: 364 NTAKVTVGKRPVTGMRLFLEGMKCNRLAIHLQYLLNTPTMLNNKIEDTTTWSEEIIDDRF 423
NT V VG RPV G+RL LEG NRLAIHLQ+L + P L+ S + +
Sbjct: 2 NTIPVDVGNRPVVGLRLQLEGRSSNRLAIHLQHLASLPKSLSVSDNSNAYLSCDSYNCNL 181
Query: 424 LEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSN 483
+ + S+VCTAPV+ + S S IVTG QL V+ L LRL FSKV
Sbjct: 182 HKKVKWNSLSYVCTAPVESDDSVS-------IVTGAQLQVEN----KCLFLRLCFSKVIG 328
Query: 484 AIVVKS-NWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPVP 542
+ K+ W Q + G S G + + + + +D KP V + SS++ P
Sbjct: 329 VTLRKAPEWDQSPSL--GQFSIKGGGILTTFISKAEQRDHPKPGDVTIGSSIYSAARLAP 502
Query: 543 VQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
V+ KLL+FVDT+++ +GP D+PGHW+V+GARL + KI L K+SL +
Sbjct: 503 VRTPKLLRFVDTTEIMRGPVDTPGHWVVSGARLHVQDAKIYLHVKYSLFS 652
>BM953913
Length = 421
Score = 138 bits (347), Expect = 8e-33
Identities = 63/140 (45%), Positives = 95/140 (67%)
Frame = +2
Query: 266 NHSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQ 325
NHS+WL T+ + PD ++ +F P T LL P + L+ AI LYL YKPP+ +L YFL++Q
Sbjct: 2 NHSKWLSTIKSSPDIIEMTFCPITDLLDEIPAKEHLTRAIGLYLEYKPPIEELRYFLEFQ 181
Query: 326 AHRLWAPIHNDLPLGPVSNRTTHSPSLSLNLMGPKLYVNTAKVTVGKRPVTGMRLFLEGM 385
+WAP+ + +P + PSL ++MG KLYV+ ++TVG+RPVTG+ L LEG
Sbjct: 182 IPCVWAPLQDKIP--GQQRKEPVCPSLQFSIMGQKLYVSQEQITVGRRPVTGLHLCLEGS 355
Query: 386 KCNRLAIHLQYLLNTPTMLN 405
K NRL++H+Q+L++ P +L+
Sbjct: 356 KQNRLSVHVQHLVSLPKILH 415
>TC235110 similar to PIR|C86410|C86410 protein F3M18.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(17%)
Length = 754
Score = 122 bits (307), Expect = 3e-28
Identities = 71/160 (44%), Positives = 104/160 (64%)
Frame = +3
Query: 432 FSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNW 491
FSHV TAPV+Y+ S S D+ A IVT VK + VL LRL FS V++A + +S W
Sbjct: 15 FSHVYTAPVQYSSS-SMDESTA-IVTKAWFEVKLVGMKKVLFLRLGFSTVASATIRRSEW 188
Query: 492 TQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPVPVQAQKLLKF 551
ST SG S + + + +S +Q DQK P V ++S+++ GPPVP + K+L F
Sbjct: 189 DGPSTTSRKSGFFS-ALMSTRLSKELQSPDQK-PNKVDINSAIYNVGPPVPTRVPKMLSF 362
Query: 552 VDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLL 591
VDT ++ +GP+D+PG+W+VTGA+L ++ G+I + AK+SLL
Sbjct: 363 VDTKEMVRGPEDTPGYWVVTGAKLCVEGGRISIKAKYSLL 482
>TC217541
Length = 746
Score = 115 bits (289), Expect = 4e-26
Identities = 73/174 (41%), Positives = 97/174 (54%), Gaps = 1/174 (0%)
Frame = +3
Query: 420 DDRFLEAISGKKFSHVCTAPVKYNPSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFS 479
D R+ E I K FSHV TAP++Y + D IVTG QL V +++VLHL+LLFS
Sbjct: 72 DSRWFEPIKWKNFSHVSTAPIEYTETSIGDLSGVHIVTGAQLGVWDFGAKNVLHLKLLFS 251
Query: 480 KVSNAIVVKSNWTQG-STQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTG 538
KV + +S W S + G+ SS+ S +D+K+ DSS+
Sbjct: 252 KVPGCTIRRSVWDHNPSAPVAQRPDGASSSLMKKTS-----EDKKE------DSSI---- 386
Query: 539 PPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
KL K VD +++ KGPQD PGHWLVTGA+L ++KGKI L K+SLLN
Sbjct: 387 -----HIGKLAKIVDMTEMSKGPQDIPGHWLVTGAKLGVEKGKIVLRIKYSLLN 533
>TC220973
Length = 986
Score = 111 bits (277), Expect = 1e-24
Identities = 70/211 (33%), Positives = 108/211 (51%), Gaps = 8/211 (3%)
Frame = +2
Query: 390 LAIHLQYLLNTPTMLNNKIEDTTT-----WS-EEIIDDRFLEAISGKKFSHVCTAPVKYN 443
L++H+Q+L++ P +L+ + W E D R+ E + K FSHV TAP++
Sbjct: 2 LSVHVQHLVSLPKILHPYWDSHVAIGAPKWQGPEEQDSRWFEPVKWKNFSHVSTAPIENP 181
Query: 444 PSWSSDKDVAFIVTGVQLHVKKHESRSVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGS 503
++ D +IVTG QL V SR+VL+++LL+S++ + +S W
Sbjct: 182 ETFIGDFSGVYIVTGAQLGVWDFGSRNVLYMKLLYSRLPGCTIRRSLWDH---------- 331
Query: 504 GSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPTGPPV--PVQAQKLLKFVDTSQLCKGP 561
K P V +++ P + A KL+K+VD S++ KGP
Sbjct: 332 ----------------VPNKPPKTVNAENTSNPDNSTLRENATANKLVKYVDLSKMTKGP 463
Query: 562 QDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
QD PGHWLVTG +L ++KGK+ L K+SLLN
Sbjct: 464 QDPPGHWLVTGGKLGVEKGKVVLRVKYSLLN 556
>CA850828 weakly similar to GP|14209545|dbj P0481E12.26 {Oryza sativa
(japonica cultivar-group)}, partial (8%)
Length = 584
Score = 109 bits (272), Expect = 4e-24
Identities = 56/123 (45%), Positives = 79/123 (63%)
Frame = +2
Query: 470 SVLHLRLLFSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVV 529
+VLHLRLLF+ + N + KS WT S + S+ FS G ++ P
Sbjct: 68 NVLHLRLLFTHIPNCSIRKSEWTAAPEASRKSFLTNLSTTFSFTQHGTTGPPKQAPT--A 241
Query: 530 VDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFS 589
++S V+P GPPVPV+A KLLK+V+T+++ +GP D+PGHWLVT A+L+ D GKI L KF+
Sbjct: 242 LNSGVYPDGPPVPVRAGKLLKYVETAEVVRGPHDAPGHWLVTAAKLVTDGGKIGLQXKFA 421
Query: 590 LLN 592
LL+
Sbjct: 422 LLD 430
>TC231359
Length = 683
Score = 95.5 bits (236), Expect = 6e-20
Identities = 49/115 (42%), Positives = 74/115 (63%)
Frame = +2
Query: 478 FSKVSNAIVVKSNWTQGSTQRSGSGSGSGSSVFSAISTSIQGKDQKKPALVVVDSSVFPT 537
++ + N + KS W G + S +++ + S + Q QK+ A V+DS V+P
Sbjct: 2 YTHIPNCSIRKSEW--GGAPEASRKSSFLTNLSTTFSFTQQSPPQKQ-APTVLDSGVYPD 172
Query: 538 GPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKICLWAKFSLLN 592
GPPVPV++ K+LK+VDTS+ +GP D+PGHWLVT A+L+ + GKI L KF+LL+
Sbjct: 173 GPPVPVRSGKMLKYVDTSEFVRGPHDAPGHWLVTAAKLVTEGGKIGLQVKFALLD 337
>AW760549
Length = 296
Score = 85.5 bits (210), Expect = 6e-17
Identities = 37/74 (50%), Positives = 51/74 (68%)
Frame = +3
Query: 267 HSEWLLTVPNKPDAVDFSFIPFTSLLKAAPGRGFLSHAINLYLRYKPPLSDLPYFLDYQA 326
HSEWL T+ ++PD + +P TSL + GF+SHAINLY RYKPP+ DL FL++Q
Sbjct: 24 HSEWLDTIDSEPDVISMLLLPLTSLWNRSGRNGFVSHAINLYHRYKPPIEDLHQFLEFQL 203
Query: 327 HRLWAPIHNDLPLG 340
R WAP+ +++ LG
Sbjct: 204 PRHWAPVASEISLG 245
>BF597072
Length = 316
Score = 73.2 bits (178), Expect = 3e-13
Identities = 38/96 (39%), Positives = 56/96 (57%)
Frame = +2
Query: 9 ALNSLGKGFDLTSDFRLKFCKGEERLVILNETERRELTVPGFGPVTDVSVDIKCDKGDLT 68
A+NS+G GFD+T D L CK RL+ +NE + R L +PG + +V IKC +G+
Sbjct: 23 AINSIGLGFDITLDITLDNCKKGARLIFVNEEQCRHLEIPGGVAIPNVPNSIKCVRGESI 202
Query: 69 RYQSDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFG 104
R +SD+L+ QM E F+Q + + SG+ FG
Sbjct: 203 RLESDVLTQYQMMEHFDQ*MLLSRNLASGHLCASFG 310
>TC230256
Length = 639
Score = 72.8 bits (177), Expect = 4e-13
Identities = 44/130 (33%), Positives = 65/130 (49%), Gaps = 5/130 (3%)
Frame = +1
Query: 468 SRSVLHLRLLFSKVSNAIVVKSNWTQ-----GSTQRSGSGSGSGSSVFSAISTSIQGKDQ 522
SR+VL+++LL+S++ + +S W T +G+ S +S +T
Sbjct: 7 SRNVLYMKLLYSRLPGCTIRRSLWDHIPNKPPKTVNAGNTSNLDNSTLKENATG------ 168
Query: 523 KKPALVVVDSSVFPTGPPVPVQAQKLLKFVDTSQLCKGPQDSPGHWLVTGARLMLDKGKI 582
KL+K+VD S++ KGPQD PGHWLVTG +L ++KGKI
Sbjct: 169 -----------------------NKLVKYVDLSEMTKGPQDPPGHWLVTGGKLGVEKGKI 279
Query: 583 CLWAKFSLLN 592
L K+SLLN
Sbjct: 280 VLRVKYSLLN 309
>BM527909 weakly similar to PIR|C86410|C864 protein F3M18.18 [imported] -
Arabidopsis thaliana, partial (7%)
Length = 427
Score = 72.0 bits (175), Expect = 7e-13
Identities = 40/89 (44%), Positives = 53/89 (58%)
Frame = +1
Query: 72 SDILSFTQMSEMFNQKSSIPGKIPSGYFNTVFGFEAGSWASDAANTKFLGLDGYVIKLFN 131
S++L+ QM E FNQ+ + G+ SG+F FG + D A+ K L DG+ IK +
Sbjct: 7 SEVLTLQQMLEHFNQEMCLGGQTASGHFCASFGLSCRN-IKDLASIKSLAYDGWFIKRYA 183
Query: 132 VHIDRYPLVLSKQVIEAVPSSWDPTALAR 160
V ++RY L V EAVPSSWDP ALAR
Sbjct: 184 VELERYQGELLDHVKEAVPSSWDPEALAR 270
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 26,865,084
Number of Sequences: 63676
Number of extensions: 392145
Number of successful extensions: 1902
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 1827
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1872
length of query: 595
length of database: 12,639,632
effective HSP length: 103
effective length of query: 492
effective length of database: 6,081,004
effective search space: 2991853968
effective search space used: 2991853968
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Lotus: description of TM0069.5


