

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0060.4
(1482 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
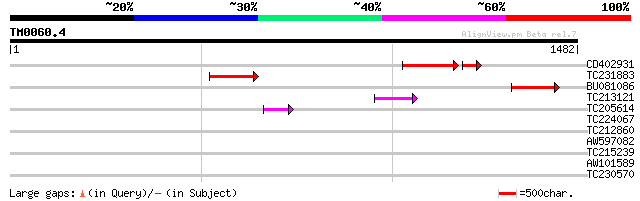
Score E
Sequences producing significant alignments: (bits) Value
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 149 2e-40
TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment... 142 1e-33
BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza s... 130 3e-30
TC213121 84 6e-16
TC205614 54 5e-07
TC224067 38 0.029
TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and... 36 0.14
AW597082 similar to GP|21666452|g polyprotein 1 {Bean pod mottle... 33 1.2
TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%) 31 3.6
AW101589 GP|21104742|db OJ1117_G01.13 {Oryza sativa (japonica cu... 30 6.1
TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (A... 30 7.9
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 149 bits (377), Expect(2) = 2e-40
Identities = 83/146 (56%), Positives = 105/146 (71%)
Frame = +2
Query: 1028 GTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHGGRTAHSLFCIPLNADEDSCCGIV 1087
GT KTFLW TL+ +RS K ++ + +ASLLL G TAHS F IPL +DS C I
Sbjct: 11 GTSKTFLWKTLSAGMRS-KGLMSSSCLKRLASLLLPRG*TAHSTFSIPLVIKDDSTCNIN 187
Query: 1088 QGSPKAELLKLSSLIIWDEAPMVSRYAFEALDRTLRDIMRICNPECFNKPFGGKVVVLGG 1147
QGS +++LL + LIIWDE P+++++ FEALDRTL+DIM N + KPFGGK VVLGG
Sbjct: 188 QGSARSKLLLHTKLIIWDETPVMNKFCFEALDRTLQDIMASQNKDNATKPFGGK-VVLGG 364
Query: 1148 DFRQILPVIPKGSRAEIVMATINSSR 1173
DFRQILPVI KGSR +IV + IN+S+
Sbjct: 365 DFRQILPVIRKGSRQDIVGSAINASK 442
Score = 36.2 bits (82), Expect(2) = 2e-40
Identities = 18/49 (36%), Positives = 30/49 (60%)
Frame = +1
Query: 1185 ENMRLFSNSETSDVEKIKVFAEWVLDIGDGVLGDYNDGDADIKVPKDTL 1233
+NMRL + + D + I+ F +W+L I DG + ++G+ DI PK+ L
Sbjct: 460 KNMRLGTTNNNEDRDDIREFVDWILMIRDGNRDEEDEGEIDI--PKNLL 600
>TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment), partial
(17%)
Length = 1263
Score = 142 bits (357), Expect = 1e-33
Identities = 67/128 (52%), Positives = 91/128 (70%)
Frame = +1
Query: 522 MYTIEFQKRGLPHAHILLWLSTKDKLHSIELIDSVICAELPDPKVYPLLYQCVSNYMVHG 581
M+TIEFQKRGLPH +LL+L ++ S + ID +I AE+P + P LY V N+MVHG
Sbjct: 877 MHTIEFQKRGLPHIQLLLFLHPDNQYPSSDEIDHIISAEIPSHEDDPELYTLVQNHMVHG 1056
Query: 582 PCGVNRTNSPCMKNGRCSKFFPKNFVEKTSFDSDGYPVYRRRNTGISTNRRGVDLDNGFV 641
PCG+ +++SPCMK G+CS+F+PK F+ T D +GY VYRRRN G + + V +DN ++
Sbjct: 1057 PCGILQSHSPCMKEGKCSRFYPKIFLPNTLLDLNGYLVYRRRNDGRTILKNSVIVDNRYL 1236
Query: 642 VPYNPKLL 649
VPYN KLL
Sbjct: 1237 VPYNAKLL 1260
>BU081086 weakly similar to GP|27817864|db OJ1081_B12.13 {Oryza sativa
(japonica cultivar-group)}, partial (7%)
Length = 426
Score = 130 bits (328), Expect = 3e-30
Identities = 60/124 (48%), Positives = 87/124 (69%)
Frame = +2
Query: 1313 ITTEFLNDIRCSGIPDHKLVLKEGAPVMLMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVI 1372
ITTEF+N + S +P+HK+ LK + +ML+RNLD + GLCN TR+++T +++ ++
Sbjct: 53 ITTEFINSLSTSRLPNHKIKLKVDSRIMLLRNLDQNEGLCNSTRLIITMFADHIIEAKIM 232
Query: 1373 SGTHVGRQVFIGRMDLMPTDGSMPIKFQRRQFPLLLSFAMTINKSQGKTLSHVGLYLPNP 1432
SG G V+I R+ P+ P K RR+FP+++S+AMTINKSQG+ L+ VGLYLP P
Sbjct: 233 SGKGQGNTVYIPRLATSPSQSPWPFKLIRRKFPIIVSYAMTINKSQGQLLASVGLYLPTP 412
Query: 1433 VFSH 1436
VFSH
Sbjct: 413 VFSH 424
>TC213121
Length = 713
Score = 83.6 bits (205), Expect = 6e-16
Identities = 44/112 (39%), Positives = 65/112 (57%)
Frame = +2
Query: 953 SLKDFDNMPCVDSNILIQYGNILLFNELNFDTVEMSKLHVECLNKLNGGQAKIYEEIISA 1012
S ++F M I+ YGN L++NEL +D ++ C + L A I+ +I+
Sbjct: 275 SHRNFPLMSYPIGYIVNPYGNKLIYNELAYDKDILAA*FSRCYHSLTDEHASIFNKIMHM 454
Query: 1013 VNSDGGEFLFVYGYGGTGKTFLWTTLTYKLRSEKKIILNVASSGIASLLLHG 1064
+ + G F+YGYGGTGKTF+W TL+ + S I+L +ASSGI S+LL G
Sbjct: 455 IATQSGGVYFLYGYGGTGKTFVWKTLSSTIHSNSGIVLTMASSGILSMLLPG 610
Score = 38.5 bits (88), Expect = 0.022
Identities = 25/77 (32%), Positives = 40/77 (51%)
Frame = +3
Query: 707 CRYLSPCEAVWRTFSFYIHDKWPPVLKQSYHLPKKQVILFNERAPIDDVLERNKVKKSMF 766
C Y+SP E + +F +H + P V +HL + E I +L ++ +K+S+F
Sbjct: 30 CIYVSPPETC*KISAFPMHGRAPTVECLYFHLENQ*P--*KESQQIGSMLSKSTIKESIF 203
Query: 767 LAWMEANCKYPLGRGLT 783
A M++N Y GR LT
Sbjct: 204 TACMDSNKMYHHGRDLT 254
>TC205614
Length = 742
Score = 53.9 bits (128), Expect = 5e-07
Identities = 29/81 (35%), Positives = 46/81 (55%), Gaps = 2/81 (2%)
Frame = +2
Query: 664 SNCIKYLFKYINKGVDRV-TMSMSVTQSSNEETTVVDEIKQYH-DCRYLSPCEAVWRTFS 721
SN ++F YI+ + S+ + ++ E + V Y+ DC+Y+SPCEA WR FS
Sbjct: 497 SNLFSFIFHYISMVENHN*RTSLKLKDLASIEASQVSFPHMYYLDCQYISPCEACWRIFS 676
Query: 722 FYIHDKWPPVLKQSYHLPKKQ 742
F I+ + P V + +HLP +Q
Sbjct: 677 FQIYGRIPIVERLYFHLPNEQ 739
>TC224067
Length = 415
Score = 38.1 bits (87), Expect = 0.029
Identities = 30/68 (44%), Positives = 41/68 (60%), Gaps = 4/68 (5%)
Frame = -2
Query: 1407 LLSFAMTINKSQGKTLSHV-GLYLPNPV-FSHGQLYVAISRVKT--RAGLKILICNEDIS 1462
L+ + M INKS G+TLS V G+ LP P SHGQ YV +++V + R K + C +
Sbjct: 363 LVCYTMKINKSHGQTLSQVGGVLLPRPN**SHGQ-YVTLNQV*SY*RRKSKSINCEKYDL 187
Query: 1463 QRDVTKNI 1470
QR +KNI
Sbjct: 186 QRIFSKNI 163
>TC212860 weakly similar to UP|PIF1_YEAST (P07271) DNA repair and recombination
protein PIF1, mitochondrial precursor, partial (8%)
Length = 596
Score = 35.8 bits (81), Expect = 0.14
Identities = 21/50 (42%), Positives = 30/50 (60%)
Frame = +2
Query: 1409 SFAMTINKSQGKTLSHVGLYLPNPVFSHGQLYVAISRVKTRAGLKILICN 1458
++AM+I+K QG TL V L F G +YVA+SRV++ GL + N
Sbjct: 2 AWAMSIHKCQGMTLERVHTDLSR-AFGCGMVYVALSRVRSLEGLHLSAFN 148
>AW597082 similar to GP|21666452|g polyprotein 1 {Bean pod mottle virus},
partial (9%)
Length = 523
Score = 32.7 bits (73), Expect = 1.2
Identities = 22/66 (33%), Positives = 35/66 (52%), Gaps = 1/66 (1%)
Frame = -3
Query: 56 FAFVEAESQMYFDLGEMNMACQYCGAILWYHERAQKAKNAISPD-FSICCMKGKISLPYL 114
FA ++ E ++ F +GE+ +AC Y IL H R + K PD ++ + I P L
Sbjct: 317 FAAIQYERELSFSIGELPVACAY--TILVTHSRD*QDKLLAFPDYYTQHNVVVSIDSPCL 144
Query: 115 QDAPTL 120
Q+ PT+
Sbjct: 143 QEPPTI 126
>TC215239 UP|H3_PEA (P02300) Histone H3, partial (39%)
Length = 340
Score = 31.2 bits (69), Expect = 3.6
Identities = 14/22 (63%), Positives = 18/22 (81%)
Frame = -3
Query: 1400 QRRQFPLLLSFAMTINKSQGKT 1421
QR FPL++ FAMT NKS+G+T
Sbjct: 338 QR**FPLIVCFAMTTNKSEGQT 273
>AW101589 GP|21104742|db OJ1117_G01.13 {Oryza sativa (japonica
cultivar-group)}, partial (24%)
Length = 235
Score = 30.4 bits (67), Expect = 6.1
Identities = 14/30 (46%), Positives = 19/30 (62%)
Frame = +1
Query: 795 KKKKEWHPRKKGICIGRMNFVPPGSGELYY 824
KKKK+ +K+G R F PPG G+LY+
Sbjct: 133 KKKKKKKKKKRGGPF*RTQFSPPGVGKLYF 222
>TC230570 similar to UP|MLO6_ARATH (Q94KB7) MLO-like protein 6 (AtMlo6),
partial (24%)
Length = 957
Score = 30.0 bits (66), Expect = 7.9
Identities = 31/100 (31%), Positives = 46/100 (46%), Gaps = 3/100 (3%)
Frame = +3
Query: 1279 NQYVLSLFPGNERTYLSSYSVWSVTEDVGIEADWITTEFLNDIRCSGI--PDHKLVLKEG 1336
N + L+ F + + S TEDV I IT L I CS + P + LV + G
Sbjct: 93 NAFQLAFFAWSALQFGIKSCFHSHTEDVVIR---ITMGVLIQILCSYVTLPLYALVTQMG 263
Query: 1337 APVM-LMRNLDVSTGLCNGTRIVVTHLRPNVVGGIVISGT 1375
+ + + N V+ L N + H+R N VG + +SGT
Sbjct: 264 STMKPTIFNDRVAVALRNWHQTAKKHIRQNRVGPLSLSGT 383
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,811,213
Number of Sequences: 63676
Number of extensions: 1017294
Number of successful extensions: 4461
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 4423
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4458
length of query: 1482
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1373
effective length of database: 5,698,948
effective search space: 7824655604
effective search space used: 7824655604
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 65 (29.6 bits)
Lotus: description of TM0060.4


