

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= TM0059.7
(1706 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
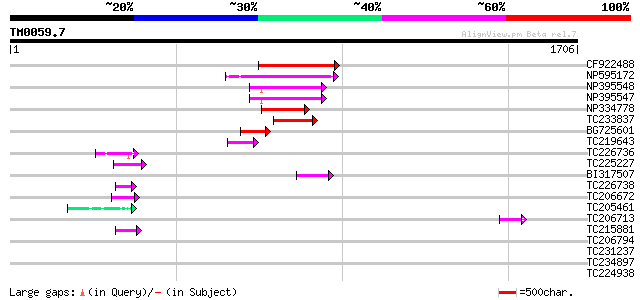
Score E
Sequences producing significant alignments: (bits) Value
CF922488 215 2e-55
NP595172 polyprotein [Glycine max] 189 9e-48
NP395548 reverse transcriptase [Glycine max] 164 3e-40
NP395547 reverse transcriptase [Glycine max] 161 3e-39
NP334778 reverse transcriptase [Glycine max] 160 6e-39
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 148 2e-35
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 97 6e-20
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 59 1e-08
TC226736 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich prot... 51 5e-06
TC225227 similar to GB|AAF02830.1|6056366|AC009894 phosphoglycer... 48 3e-05
BI317507 46 2e-04
TC226738 weakly similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-ri... 44 8e-04
TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyra... 44 8e-04
TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MS... 44 8e-04
TC206713 similar to GB|AAT41855.1|48310625|BT014872 At2g19340 {A... 44 8e-04
TC215881 homologue to UP|Q8CBU4 (Q8CBU4) Mus musculus adult male... 44 8e-04
TC206794 weakly similar to UP|Q6UAB9 (Q6UAB9) SufC, partial (75%) 43 0.001
TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644 At2g4... 43 0.001
TC234897 weakly similar to UP|TIR3_YEAST (P40552) Cell wall prot... 43 0.001
TC224938 similar to GB|AAS75312.1|45680886|AY568529 multidomain ... 42 0.002
>CF922488
Length = 741
Score = 215 bits (547), Expect = 2e-55
Identities = 111/242 (45%), Positives = 153/242 (62%)
Frame = +3
Query: 750 VKKANGKWRMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHR 809
V K +GK MCVDY DLN A PKD FPLP I+ LVDN++ + S MD +SGYNQI +
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 810 DDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPK 869
+D EKT FIT GT+CY + FGLKN ATYQR M +F D+M K +EVY+DD+IVK+
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 870 GGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMK 929
+H +L +F +L+K+ +RLNP KC F V+S K L ++ + RGIE++ +K + I+ M
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 930 SPRTVKEVQQLAGRMAAIGRFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEILS 989
P T K+VQ GR+ I RF+ PL+ LL +W + AF ++K+ L
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 990 SP 991
+P
Sbjct: 723 NP 728
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 189 bits (480), Expect = 9e-48
Identities = 114/341 (33%), Positives = 185/341 (53%)
Frame = +1
Query: 649 LPENTTNLARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSVDRKYKPVA 708
+PE+T + LP ++ + LL + ++FA + P + H + + + PV
Sbjct: 1636 VPEDTL---KDLPTNIDPELAILLHTYAQVFAVPASLPPQREQ---DHAIPLKQGSGPVK 1797
Query: 709 QKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANGKWRMCVDYTDLNK 768
+ + ++ I++ E+L GII+ L ++LVKK +G WR C DY LN
Sbjct: 1798 VRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSPFSLP-ILLVKKKDGSWRFCTDYRALNA 1974
Query: 769 ACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTM 828
KD FP+P++D L+D G +Y S +D SGY+QI + +D EKTAF T G Y + +
Sbjct: 1975 ITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEWLV 2154
Query: 829 LPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQLKKHN 888
+PFGL NA AT+Q +M +IF + K V V+ DDI++ + DH L V + LK+H
Sbjct: 2155 MPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQ 2334
Query: 889 MRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQLAGRMAAIG 948
+ KC+FG +LG+ ++ G+ + K QA+++ +P VK+++ G
Sbjct: 2335 LFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYR 2514
Query: 949 RFLPKAALRALPLYTLLKNGATFEWSAEADAAFTQLKEILS 989
RF+ A A PL LL+ +F W+ EA+AAF +LK+ ++
Sbjct: 2515 RFIKSYANIAGPLTDLLQKD-SFLWNNEAEAAFVKLKKAMT 2634
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 164 bits (415), Expect = 3e-40
Identities = 86/248 (34%), Positives = 134/248 (53%), Gaps = 18/248 (7%)
Frame = +1
Query: 722 IQQQTTELLRAGIIREVKYTTWLSNVVLVKKANG------------------KWRMCVDY 763
++++ +LL G+I + + W+S V++V K G W++C+DY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 764 TDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGT 823
LN+A KD FPLP +D +++ +G+ Y +DAY GYNQI + D+EK AF G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 824 YCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQ 883
+ Y +PFGL NA T+Q M IF D++ KS+EV++DD V P L +V ++
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 884 LKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQLAGR 943
+ N+ LN +KC F VR G LG+ ++ RGIE++ K I + P VK ++ G+
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 944 MAAIGRFL 951
RF+
Sbjct: 721 ARFYRRFI 744
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 161 bits (407), Expect = 3e-39
Identities = 86/248 (34%), Positives = 134/248 (53%), Gaps = 18/248 (7%)
Frame = +1
Query: 722 IQQQTTELLRAGIIREVKYTTWLSNVVLVKKANG------------------KWRMCVDY 763
++++ +LL AG+I + ++W+S V +V K G +WRMC+DY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 764 TDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAFITDRGT 823
LN+A KD +PLP +D ++ + + +D YSGYNQI + D+EKTAF
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 824 YCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADLAVVFEQ 883
+ Y +PFGL NA T+QR M IF D++ K +EV++DD G+ A+L V ++
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 884 LKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDKCQAIMNMKSPRTVKEVQQLAGR 943
+K N+ LN +KC F V+ G LG+ ++ RGIE+ +K I + P VK + G
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 944 MAAIGRFL 951
+ RF+
Sbjct: 721 VGFYRRFI 744
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 160 bits (404), Expect = 6e-39
Identities = 75/143 (52%), Positives = 98/143 (68%)
Frame = +3
Query: 758 RMCVDYTDLNKACPKDPFPLPSIDALVDNSSGYEYLSLMDAYSGYNQIPMHRDDEEKTAF 817
RMCVDY DLN+A PKD FPLP ID L+ N + + S MD +SGYNQI M +D EKT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 818 ITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMGKSVEVYIDDIIVKTPKGGDHAADL 877
IT GT+CY ++ FGLKN ATY R M +F D+M K +E Y+D++I K+ +H +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 878 AVVFEQLKKHNMRLNPDKCTFGV 900
+F QL+K+ +RLNP KC FG+
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 148 bits (374), Expect = 2e-35
Identities = 67/132 (50%), Positives = 98/132 (73%)
Frame = +2
Query: 794 SLMDAYSGYNQIPMHRDDEEKTAFITDRGTYCYTMLPFGLKNAVATYQRMMTRIFGDLMG 853
S MD +SGYNQI M R+D EKT F+T GT+ Y ++ FGLKN ATYQR M +F D+M
Sbjct: 5 SFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMMH 184
Query: 854 KSVEVYIDDIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNR 913
K +EVY+DD+I K+ +H +L +F +L+K+ ++LNP KCTFGV+SGK LG++++ +
Sbjct: 185 KEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQK 364
Query: 914 GIELNPDKCQAI 925
GIE++P+K +A+
Sbjct: 365 GIEIDPEKVKAL 400
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 97.1 bits (240), Expect = 6e-20
Identities = 44/90 (48%), Positives = 63/90 (69%)
Frame = -3
Query: 696 HRLSVDRKYKPVAQKKRQMSAEKQQVIQQQTTELLRAGIIREVKYTTWLSNVVLVKKANG 755
H+L++ K V Q+KR++ E+ Q + Q+ +L A IR++ Y+T L +VV+VKK NG
Sbjct: 277 HKLAICNDVKLVTQRKRKIREERCQTV*QEVVKLAIASFIRDINYST*LFSVVMVKKPNG 98
Query: 756 KWRMCVDYTDLNKACPKDPFPLPSIDALVD 785
KWR+C DY DLN ACPKD +PLP+ID + D
Sbjct: 97 KWRICTDYIDLN*ACPKDAYPLPNIDHMTD 8
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 59.3 bits (142), Expect = 1e-08
Identities = 28/93 (30%), Positives = 49/93 (52%)
Frame = +3
Query: 656 LARGLPRDLSKRMEKLLLSNGKLFAWSSEDMPGIDPAFCSHRLSVDRKYKPVAQKKRQMS 715
+ G+ + + + LL +FAWS +DMPG+ HRL ++ + PV QK R+M
Sbjct: 1032 IGTGITAPIREELIILLRDYQDIFAWSYQDMPGLSSDIVQHRLPLNPECSPVKQKLRRMK 1211
Query: 716 AEKQQVIQQQTTELLRAGIIREVKYTTWLSNVV 748
E I+++ + AG + +Y W++N+V
Sbjct: 1212 PETSLKIKEEVKK*FDAGFLAVARYPKWVANIV 1310
>TC226736 similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein, partial
(63%)
Length = 1243
Score = 50.8 bits (120), Expect = 5e-06
Identities = 55/155 (35%), Positives = 69/155 (44%), Gaps = 23/155 (14%)
Frame = +2
Query: 257 RDRPPPLLTRGDKLDSKRFCEFH-DSPGHNTDECLNLKDKVEELIRAGRLS----RYVAV 311
R +PPP R + + DSP D ++ +V+ R G + R V+
Sbjct: 356 RRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADS--PVRRRVDSPYRRGGDTPPRRRPVSP 529
Query: 312 STGALPRP----RSP----PPRRTPTPPRHRDQTPP-KRRSAERRDKTPPRR-------- 354
G P P RSP P R +P R R PP +RRS RR ++PPRR
Sbjct: 530 GRGRSPSPPRRLRSPARLSPRRMRGSPGRRRSPPPPPRRRSPPRRPRSPPRRSPIGRRRT 709
Query: 355 RSPERR-RRSRERRRSPDRHGRDEVRRQHGSNIVD 388
RSP RR RSR R SP R GR VRR S+ D
Sbjct: 710 RSPIRRSARSRSRSFSP-RRGRPPVRRGRSSSYSD 811
Score = 38.5 bits (88), Expect = 0.026
Identities = 23/55 (41%), Positives = 26/55 (46%), Gaps = 1/55 (1%)
Frame = +2
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRR-SPERRRRSRERRRSPD 371
R R PRR P P R P+R + RR +P RR SP RRR RR D
Sbjct: 335 RQRESSPRRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADSPVRRRVDSPYRRGGD 499
Score = 37.4 bits (85), Expect = 0.057
Identities = 31/79 (39%), Positives = 40/79 (50%), Gaps = 11/79 (13%)
Frame = +2
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAER-RDKTPPRRRSPERRRRS----------RER 366
RPRSPP RR+P R R ++P +R + R R +P R R P RR RS +
Sbjct: 662 RPRSPP-RRSPI-GRRRTRSPIRRSARSRSRSFSPRRGRPPVRRGRSSSYSDSPSPRKVS 835
Query: 367 RRSPDRHGRDEVRRQHGSN 385
RRS R R +R + SN
Sbjct: 836 RRSKSRSPRRPLRGRGSSN 892
Score = 36.6 bits (83), Expect = 0.097
Identities = 24/61 (39%), Positives = 29/61 (47%), Gaps = 2/61 (3%)
Frame = +2
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRR--SPERRRRSRERRRSPDRHG 374
PR + PP R +P R +P + S RR +P RRR SP RR RR P G
Sbjct: 353 PRRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADSPVRRRVDSPYRRGGDTPPRRRPVSPG 532
Query: 375 R 375
R
Sbjct: 533 R 535
Score = 35.4 bits (80), Expect = 0.22
Identities = 25/62 (40%), Positives = 30/62 (48%), Gaps = 3/62 (4%)
Frame = +2
Query: 323 PPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRS-PERRRRSRERRR--SPDRHGRDEVR 379
P R+ + PR + P+RRS R PRR P RR S RRR SP R G D
Sbjct: 329 PKRQRESSPRRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADSPVRRRVDSPYRRGGDTPP 508
Query: 380 RQ 381
R+
Sbjct: 509 RR 514
Score = 33.5 bits (75), Expect = 0.82
Identities = 28/82 (34%), Positives = 32/82 (38%), Gaps = 23/82 (28%)
Frame = +2
Query: 323 PPRRTPTPPRHRDQTPPKR--------------------RSAERRDKTP--PRRRSPERR 360
PPR+ +PP PKR R + R K P PRRRSP R
Sbjct: 224 PPRQKVSPPPKTSAVAPKRDTPRTDNASADVDKDGPKRQRESSPRRKPPPSPRRRSPVPR 403
Query: 361 RRSRERR-RSPDRHGRDEVRRQ 381
R RR SP R VRR+
Sbjct: 404 RAGSPRRPDSPRRRADSPVRRR 469
Score = 33.1 bits (74), Expect = 1.1
Identities = 25/79 (31%), Positives = 37/79 (46%), Gaps = 9/79 (11%)
Frame = +2
Query: 317 PRPR-SPPPRRTPTPPRHRDQTPPKRRSAE--------RRDKTPPRRRSPERRRRSRERR 367
PR + SPPP+ + P+ RD SA+ +R+ +P R+ P RRRS R
Sbjct: 227 PRQKVSPPPKTSAVAPK-RDTPRTDNASADVDKDGPKRQRESSPRRKPPPSPRRRSPVPR 403
Query: 368 RSPDRHGRDEVRRQHGSNI 386
R+ D RR+ S +
Sbjct: 404 RAGSPRRPDSPRRRADSPV 460
>TC225227 similar to GB|AAF02830.1|6056366|AC009894 phosphoglycerate kinase
{Arabidopsis thaliana;} , partial (35%)
Length = 612
Score = 48.1 bits (113), Expect = 3e-05
Identities = 35/105 (33%), Positives = 45/105 (42%), Gaps = 4/105 (3%)
Frame = +2
Query: 311 VSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSP 370
+S + P PR PPP P PPR PP R+ R PRR RR R RR
Sbjct: 119 LSLSSTPPPRPPPPPPPPPPPRACSSPPPSARARSRAALASPRRTRCSRRMSQRASRR*E 298
Query: 371 DRHGRDE----VRRQHGSNIVDVGSIAGGWAAGGPTNNSQKRSTR 411
+ R RR GS + G ++ GPT+ S+ +TR
Sbjct: 299 EERARGGWCRWRRRAWGS*A--RRT*RGRRSSSGPTSTSRSTTTR 427
>BI317507
Length = 359
Score = 45.8 bits (107), Expect = 2e-04
Identities = 28/113 (24%), Positives = 53/113 (46%), Gaps = 1/113 (0%)
Frame = -1
Query: 862 DIIVKTPKGGDHAADLAVVFEQLKKHNMRLNPDKCTFGVRSGKFLGYMLTNRGIELNPDK 921
+I++ +P H L V + LKK + N KC F + ++LG++++ + ++ +K
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 922 CQAIMNMKSPRTVKEVQQLAGRMAAIGRFLPK-AALRALPLYTLLKNGATFEW 973
++++ P+ VK V +F+ L PL L KN F+W
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLTKNDG-FKW 18
>TC226738 weakly similar to UP|Q9SEE9 (Q9SEE9) Arginine/serine-rich protein,
partial (24%)
Length = 486
Score = 43.5 bits (101), Expect = 8e-04
Identities = 29/67 (43%), Positives = 33/67 (48%), Gaps = 3/67 (4%)
Frame = +3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRR-SPERRRRSRERRR--SPDRHG 374
R R PRR P P R P+R + RR +P RR SP RR S RRR SP R G
Sbjct: 102 RQRESSPRRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADSPRRRADSPVRRRLDSPYRRG 281
Query: 375 RDEVRRQ 381
D R+
Sbjct: 282 GDTPPRR 302
Score = 40.4 bits (93), Expect = 0.007
Identities = 26/62 (41%), Positives = 31/62 (49%), Gaps = 2/62 (3%)
Frame = +3
Query: 321 SPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRR--SRERRRSPDRHGRDEV 378
+PP RR +P R R +PP+R + R R SP RRR RRRSP R R
Sbjct: 288 TPPRRRPVSPGRGRSPSPPRRLRSPARLSPRRIRGSPGRRRSPPPPPRRRSPPRRPRSPP 467
Query: 379 RR 380
RR
Sbjct: 468 RR 473
Score = 37.7 bits (86), Expect = 0.044
Identities = 28/80 (35%), Positives = 36/80 (45%), Gaps = 14/80 (17%)
Frame = +3
Query: 321 SPPPRRTPTPPRHR-----------DQTPPKRR--SAERRDKTP-PRRRSPERRRRSRER 366
SPPP+ + P+ D+ PKR+ S+ RR P PRRRSP RR R
Sbjct: 9 SPPPKASAVAPKRDTPRADNASADVDKDGPKRQRESSPRRKPPPSPRRRSPVPRRAGSPR 188
Query: 367 RRSPDRHGRDEVRRQHGSNI 386
R R D RR+ S +
Sbjct: 189 RPDSPRRRADSPRRRADSPV 248
Score = 37.4 bits (85), Expect = 0.057
Identities = 24/63 (38%), Positives = 29/63 (45%), Gaps = 2/63 (3%)
Frame = +3
Query: 323 PPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRR--SPDRHGRDEVRR 380
P R+ + PR + P+RRS R PRR RRR RRR SP R D R
Sbjct: 96 PKRQRESSPRRKPPPSPRRRSPVPRRAGSPRRPDSPRRRADSPRRRADSPVRRRLDSPYR 275
Query: 381 QHG 383
+ G
Sbjct: 276 RGG 284
Score = 35.4 bits (80), Expect = 0.22
Identities = 25/58 (43%), Positives = 28/58 (48%), Gaps = 3/58 (5%)
Frame = +3
Query: 317 PRPRSPP-PRRTPTPPRHRDQTPPKRRSAERRDKTPPRRR--SPERRRRSRERRRSPD 371
PR + PP PRR PR P+R + RR PRRR SP RRR RR D
Sbjct: 120 PRRKPPPSPRRRSPVPRRAGS--PRRPDSPRRRADSPRRRADSPVRRRLDSPYRRGGD 287
Score = 35.0 bits (79), Expect = 0.28
Identities = 18/37 (48%), Positives = 21/37 (56%)
Frame = +3
Query: 318 RPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRR 354
R R P RR PP PP+RRS RR ++PPRR
Sbjct: 381 RIRGSPGRRRSPPP------PPRRRSPPRRPRSPPRR 473
>TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyrate
dehydrogenase , partial (76%)
Length = 1230
Score = 43.5 bits (101), Expect = 8e-04
Identities = 30/89 (33%), Positives = 39/89 (43%), Gaps = 3/89 (3%)
Frame = +3
Query: 305 LSRYVAVSTGALPRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSR 364
L R ++ L RP PPPRR P H P +R R + P R P RRR R
Sbjct: 285 LGRPLSPPPPLLRRPLRPPPRRRPRRHDHLRTLPRRRNRGRRYRQRLPLNRRPRVRRRPR 464
Query: 365 ERRRSPD---RHGRDEVRRQHGSNIVDVG 390
R+P R GR + +GS ++ G
Sbjct: 465 REERNPSNLRRRGRID-GEAYGSLVLPFG 548
Score = 32.7 bits (73), Expect = 1.4
Identities = 20/50 (40%), Positives = 21/50 (42%)
Frame = +3
Query: 317 PRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRER 366
P P P + P PPR R P PPRRRSP RRR R
Sbjct: 159 PHRLQPHPLQGPAPPRPRGLLRP----------LPPRRRSPLRRRLLHRR 278
>TC205461 similar to GB|AAN28790.1|23505861|AY143851 At5g64200/MSJ1_4
{Arabidopsis thaliana;} , partial (69%)
Length = 1148
Score = 43.5 bits (101), Expect = 8e-04
Identities = 57/213 (26%), Positives = 84/213 (38%), Gaps = 7/213 (3%)
Frame = +3
Query: 175 EFLARAEKFINMEDA--VTLRAASQTLAIKGPQKMKEHKDQPSTRESRRKGQDERKPKRK 232
++ A+K + D V R + A GP + HK + R + R P+++
Sbjct: 285 KYADEAQKAVERLDGRMVDGREITVQFAKYGPNAERIHKGRIIETSPRSRRSRSRSPRKR 464
Query: 233 KYDSYTPLNSSLSRILREKASTDLRDRPPPLLTRGDKLDSKRFCEFHDSPGHNTDECLNL 292
D Y R R ++ + DR RG D +R + L+
Sbjct: 465 HRDDYR------DRDYRRRSRSRSYDRYERDRHRGRDKDYRR-------RSRSRSASLDY 605
Query: 293 KDKVEELIRAGRLSRYVAVSTGAL-PRPRSPPPRRTPTPPRHRDQTPPKRRSAERRDKTP 351
K + R SR + S + P RSP PRR+P+P R T P+R ++ R+
Sbjct: 606 KGRGRGRYDDERNSRSRSRSVDSRSPARRSPSPRRSPSPQR---STSPQRSTSPRKS--- 767
Query: 352 PRRRSPERRRRSRERR----RSPDRHGRDEVRR 380
PR SP RSRE R RS GR + R
Sbjct: 768 PRGESP--ANRSREERSPTPRSVSPRGRPDASR 860
Score = 32.3 bits (72), Expect = 1.8
Identities = 28/101 (27%), Positives = 38/101 (36%), Gaps = 7/101 (6%)
Frame = +3
Query: 335 DQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRD-------EVRRQHGSNIV 387
+ +P RRS R + R +R R R R RS DR+ RD + RR+ S
Sbjct: 414 ETSPRSRRSRSRSPRKRHRDDYRDRDYRRRSRSRSYDRYERDRHRGRDKDYRRRSRSRSA 593
Query: 388 DVGSIAGGWAAGGPTNNSQKRSTRVIMSAAGRPRPGSLHPP 428
+ G NS+ RS V + R P P
Sbjct: 594 SLDYKGRGRGRYDDERNSRSRSRSVDSRSPARRSPSPRRSP 716
>TC206713 similar to GB|AAT41855.1|48310625|BT014872 At2g19340 {Arabidopsis
thaliana;} , partial (54%)
Length = 449
Score = 43.5 bits (101), Expect = 8e-04
Identities = 33/88 (37%), Positives = 45/88 (50%), Gaps = 5/88 (5%)
Frame = +3
Query: 1473 SSTRTSSAGSESPES*SQTMGPSLPLRGSETSWMG---YTSNIVSPPSSTPRR--TDRQS 1527
SST ++S+ S P S S PS P S TSW +T++ SPP+S P R T S
Sbjct: 156 SSTSSASSASSFPPSPSPPRTPSSPSSSSPTSWSSPALFTTSSSSPPASAPLRTPTPAPS 335
Query: 1528 QPTGSFSGASGKDWGAPRATGPNSSITS 1555
P+ S AS + RA+ P+S +S
Sbjct: 336 APSSSCPAASTASTSS-RASPPDSCSSS 416
>TC215881 homologue to UP|Q8CBU4 (Q8CBU4) Mus musculus adult male urinary
bladder cDNA, RIKEN full-length enriched library,
clone:9530007N19 product:villin 2, full insert sequence.
(Fragment), partial (5%)
Length = 1049
Score = 43.5 bits (101), Expect = 8e-04
Identities = 28/80 (35%), Positives = 35/80 (43%), Gaps = 2/80 (2%)
Frame = -2
Query: 318 RPRSPPPRRTPTPPRHRDQT--PPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGR 375
+P +PP +P PPR R + P R R + PP +P +RRS RRR GR
Sbjct: 574 KP*TPPTPTSPAPPRIRARCLGPDPPRPPPRGSRRPPAPPAPSAKRRSARRRRRSGGGGR 395
Query: 376 DEVRRQHGSNIVDVGSIAGG 395
RR GS GG
Sbjct: 394 RGARRARRRRRRRRGSAGGG 335
Score = 40.8 bits (94), Expect = 0.005
Identities = 33/78 (42%), Positives = 35/78 (44%), Gaps = 10/78 (12%)
Frame = -2
Query: 317 PRPRSPPPRRT------PTPPRHRDQTPPKRRSAERRDKTPP----RRRSPERRRRSRER 366
P P SP P R P PPR PP R S RR PP +RRS RRRRS
Sbjct: 559 PTPTSPAPPRIRARCLGPDPPR-----PPPRGS--RRPPAPPAPSAKRRSARRRRRSGGG 401
Query: 367 RRSPDRHGRDEVRRQHGS 384
R R R RR+ GS
Sbjct: 400 GRRGARRARRRRRRRRGS 347
Score = 36.2 bits (82), Expect = 0.13
Identities = 31/100 (31%), Positives = 37/100 (37%), Gaps = 3/100 (3%)
Frame = +3
Query: 312 STGALPRPRSPPPRRTPTPPRHRDQTPPKR---RSAERRDKTPPRRRSPERRRRSRERRR 368
S A R + PPPR T P R PP R R PRR P RR R RR
Sbjct: 261 SQAASRRRQRPPPRHHSTLPLSRHHPPPAEPLLRRRRLRALRAPRRPPPPLLRRRRALRR 440
Query: 369 SPDRHGRDEVRRQHGSNIVDVGSIAGGWAAGGPTNNSQKR 408
+ G RR+ GG GP + ++ R
Sbjct: 441 FAEGAGGAGGRRE---------PRGGGRGGSGPKHRARMR 533
>TC206794 weakly similar to UP|Q6UAB9 (Q6UAB9) SufC, partial (75%)
Length = 1316
Score = 43.1 bits (100), Expect = 0.001
Identities = 24/61 (39%), Positives = 32/61 (52%)
Frame = +1
Query: 322 PPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRRQ 381
PPPR+ P R +PP E +D RR SP RR+ R+ RR P HG+ R++
Sbjct: 124 PPPRQPPRLRGQRRHSPPPS**LEGQDCRIQRRHSP-RRQPHRQPRRGPRHHGKKRFRQK 300
Query: 382 H 382
H
Sbjct: 301 H 303
>TC231237 weakly similar to GB|AAL69460.1|18491271|AY074644
At2g41330/F13H10.12 {Arabidopsis thaliana;} , partial
(24%)
Length = 583
Score = 43.1 bits (100), Expect = 0.001
Identities = 34/96 (35%), Positives = 39/96 (40%), Gaps = 26/96 (27%)
Frame = +3
Query: 301 RAGRLSRYVAVSTGALPRP-----------RSPPPRRTPTPPRHRDQTPPKRRSAERRDK 349
R GRL+R T A PRP R P RR P P H + P +RR A R +
Sbjct: 99 RLGRLTRRRPRRTPARPRPGCRRLLHLPPRRPPHLRRLPRRPIHPPRPPRRRRRARRLHR 278
Query: 350 TPPRRRSPER---------------RRRSRERRRSP 370
P RR+P R RR R RRR P
Sbjct: 279 RPLPRRAPRRPRLPRQPRPAACLRRRRLRRRRRRRP 386
Score = 31.6 bits (70), Expect = 3.1
Identities = 21/52 (40%), Positives = 24/52 (45%), Gaps = 1/52 (1%)
Frame = +3
Query: 320 RSPPPRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSP-ERRRRSRERRRSP 370
R P PRR P PR Q P RR + RRR P R RR+ R+P
Sbjct: 276 RRPLPRRAPRRPRLPRQPRPAACLRRRRLRRRRRRRPPAPRERRAPPPHRTP 431
>TC234897 weakly similar to UP|TIR3_YEAST (P40552) Cell wall protein TIR3
precursor, partial (12%)
Length = 698
Score = 43.1 bits (100), Expect = 0.001
Identities = 30/67 (44%), Positives = 33/67 (48%), Gaps = 3/67 (4%)
Frame = +2
Query: 319 PRSPPPRRTPTPPRHRDQTP-PKRRSAERRDKTP--PRRRSPERRRRSRERRRSPDRHGR 375
P+ PPPRR R Q+P P RR R D P PRRR RRRR R RRR
Sbjct: 179 PQHPPPRRLHRQRRRFPQSPHPHRRLHRRHDVAPQHPRRR---RRRRRRRRRRHCPLPPS 349
Query: 376 DEVRRQH 382
D +R H
Sbjct: 350 DRQQRPH 370
Score = 35.8 bits (81), Expect = 0.17
Identities = 24/63 (38%), Positives = 27/63 (42%), Gaps = 14/63 (22%)
Frame = +2
Query: 320 RSPPPRRTPTPPRHRDQTPPKRRSAERRDKTP----PRRR----------SPERRRRSRE 365
+ P PR P P Q PP RR +R + P P RR P RRRR R
Sbjct: 134 QEPRPRPRPRPQVQAPQHPPPRRLHRQRRRFPQSPHPHRRLHRRHDVAPQHPRRRRRRRR 313
Query: 366 RRR 368
RRR
Sbjct: 314 RRR 322
Score = 35.0 bits (79), Expect = 0.28
Identities = 32/91 (35%), Positives = 35/91 (38%), Gaps = 28/91 (30%)
Frame = +2
Query: 320 RSPPPRR-TPTPPRHRDQTPPKRRSAERRDKTPPRRR--------------------SPE 358
RSPPP P P RHR RR+A R PPRRR P
Sbjct: 17 RSPPPNAPNPRPLRHR-HLHLLRRAAPLRAPHPPRRR*GLQEPRPRPRPRPQVQAPQHPP 193
Query: 359 RRRRSRERRRSPDR-------HGRDEVRRQH 382
RR R+RRR P H R +V QH
Sbjct: 194 PRRLHRQRRRFPQSPHPHRRLHRRHDVAPQH 286
>TC224938 similar to GB|AAS75312.1|45680886|AY568529 multidomain cyclophilin
type peptidyl-prolyl cis-trans isomerase {Arabidopsis
thaliana;} , partial (17%)
Length = 1474
Score = 42.0 bits (97), Expect = 0.002
Identities = 29/63 (46%), Positives = 34/63 (53%)
Frame = +2
Query: 324 PRRTPTPPRHRDQTPPKRRSAERRDKTPPRRRSPERRRRSRERRRSPDRHGRDEVRRQHG 383
PRR +PP R +TPP+ RS RR +TP RSP R R R RSP R R V R
Sbjct: 818 PRRFRSPPP-RGRTPPRYRS--RRSRTPSVSRSPRYRARRYSRSRSPVR-SRSLVDRSRP 985
Query: 384 SNI 386
S +
Sbjct: 986 SRV 994
Score = 31.6 bits (70), Expect = 3.1
Identities = 20/48 (41%), Positives = 25/48 (51%), Gaps = 1/48 (2%)
Frame = +2
Query: 334 RDQTPPKRRSAERRDKTPPRR-RSPERRRRSRERRRSPDRHGRDEVRR 380
RD+ RR + RR ++PP R R+P R R R R S R R RR
Sbjct: 785 RDRYSTYRRFSPRRFRSPPPRGRTPPRYRSRRSRTPSVSRSPRYRARR 928
Score = 30.4 bits (67), Expect = 7.0
Identities = 22/71 (30%), Positives = 33/71 (45%), Gaps = 13/71 (18%)
Frame = +2
Query: 329 TPPRHRDQTPPKRRSAERRDKTPPR-------RRSPERRRRSRERRRSPDRH------GR 375
+P R ++P + R+ +++P R RSP R R R RRSP R R
Sbjct: 23 SPSRSISRSPVRGRNGRSVNRSPVRGRNGRSVSRSPVRGRNGRSVRRSPVRGRNGRSVSR 202
Query: 376 DEVRRQHGSNI 386
VR ++G N+
Sbjct: 203 SPVRGRNGRNV 235
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.443
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 81,854,550
Number of Sequences: 63676
Number of extensions: 1382718
Number of successful extensions: 22620
Number of sequences better than 10.0: 743
Number of HSP's better than 10.0 without gapping: 14154
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 19696
length of query: 1706
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1596
effective length of database: 5,635,272
effective search space: 8993894112
effective search space used: 8993894112
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 66 (30.0 bits)
Lotus: description of TM0059.7


